This notebook turns a small PyTorch dense network into a lecture-style diagram like the hand-drawn sketch.
New capability:
- for small fully connected networks, the visualizer can annotate edge weights, node biases, and forward-pass values
- if you pass an
input_vector, it computes z and a layer by layer
It works for:
nn.Sequential(...) models made of nn.Linear + common activations- custom
nn.Module classes that still form a feed-forward fully connected network
It is intentionally not for CNNs, residual blocks, attention, or branching graphs.
import sys
from pathlib import Path
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
torch.manual_seed(7)
repo_root = Path.cwd().resolve()
if repo_root.name == "lecture_demo":
repo_root = repo_root.parent
if str(repo_root) not in sys.path:
sys.path.append(str(repo_root))
from fc_model_visualizer import (
extract_fully_connected_architecture,
save_figure,
visualize_fully_connected_model,
)
plt.rcParams["figure.figsize"] = (12, 6)
print(f"Repo root: {repo_root}")
print(f"Torch version: {torch.__version__}")
%config InlineBackend.figure_format = "retina"
Repo root: /Users/nipun/git/principles-ai-teaching
Torch version: 2.9.1
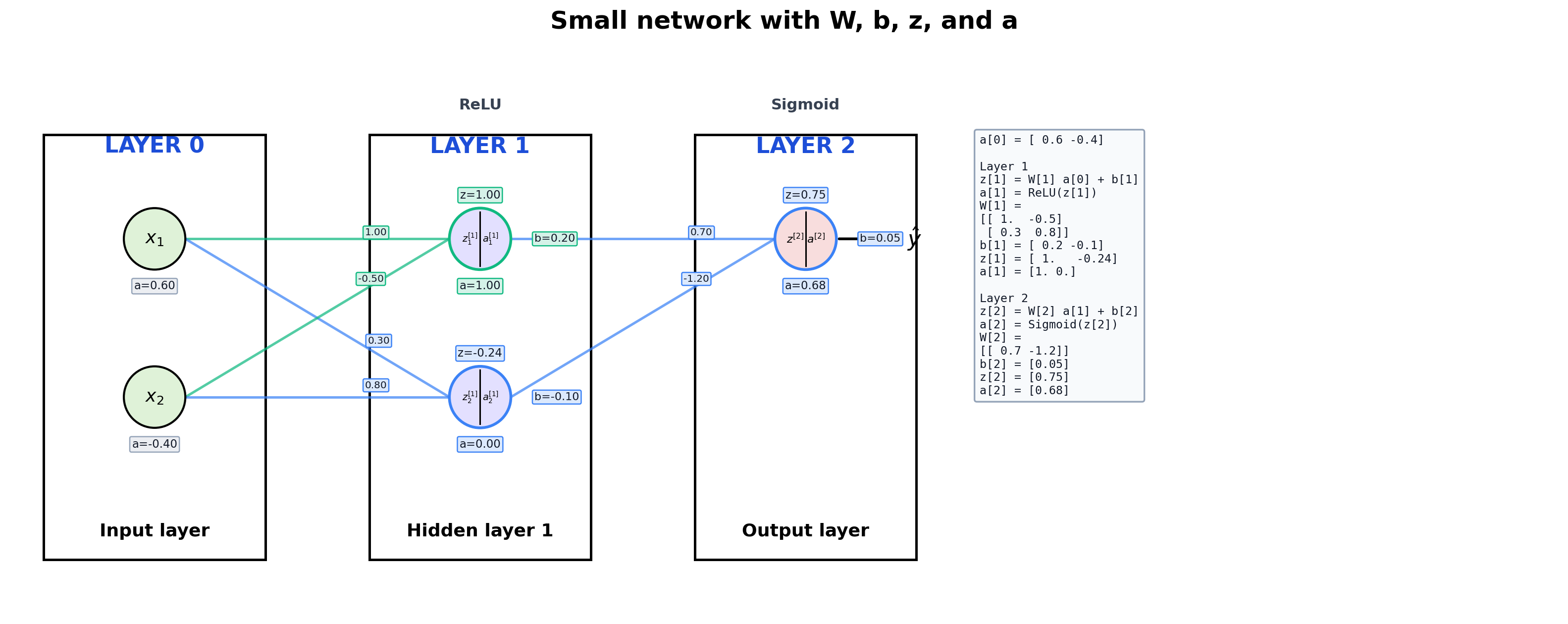
1. Small Network With Full Annotation
This is the best teaching case because we can see everything:
W on the edgesb near each neuronz = W a + ba = g(z)- the actual matrix values in a side panel
Design cue:
- incoming edges that feed the same neuron share a color
- the
z, a, and b badges for that neuron use the same accent color
small_model = nn.Sequential(
nn.Linear(2, 2),
nn.ReLU(),
nn.Linear(2, 1),
nn.Sigmoid(),
)
with torch.no_grad():
small_model[0].weight.copy_(torch.tensor([[1.0, -0.5], [0.3, 0.8]]))
small_model[0].bias.copy_(torch.tensor([0.2, -0.1]))
small_model[2].weight.copy_(torch.tensor([[0.7, -1.2]]))
small_model[2].bias.copy_(torch.tensor([0.05]))
x_example = torch.tensor([0.6, -0.4], dtype=torch.float32)
print(small_model)
print()
print("Architecture:")
print(extract_fully_connected_architecture(small_model))
print()
print("Input example:", x_example)
print("W[1] =")
print(small_model[0].weight.detach())
print("b[1] =", small_model[0].bias.detach())
print("W[2] =")
print(small_model[2].weight.detach())
print("b[2] =", small_model[2].bias.detach())
Sequential(
(0): Linear(in_features=2, out_features=2, bias=True)
(1): ReLU()
(2): Linear(in_features=2, out_features=1, bias=True)
(3): Sigmoid()
)
Architecture:
[LinearLayerSpec(in_features=2, out_features=2, activation='ReLU', name='0'), LinearLayerSpec(in_features=2, out_features=1, activation='Sigmoid', name='2')]
Input example: tensor([ 0.6000, -0.4000])
W[1] =
tensor([[ 1.0000, -0.5000],
[ 0.3000, 0.8000]])
b[1] = tensor([ 0.2000, -0.1000])
W[2] =
tensor([[ 0.7000, -1.2000]])
b[2] = tensor([0.0500])
fig, ax = visualize_fully_connected_model(
small_model,
input_labels=[r"$x_1$", r"$x_2$"],
input_vector=x_example,
title="Small network with W, b, z, and a",
max_neurons_per_layer=5,
show_edge_weights=True,
show_biases=True,
show_values=True,
show_matrix_details=True,
)
plt.show()
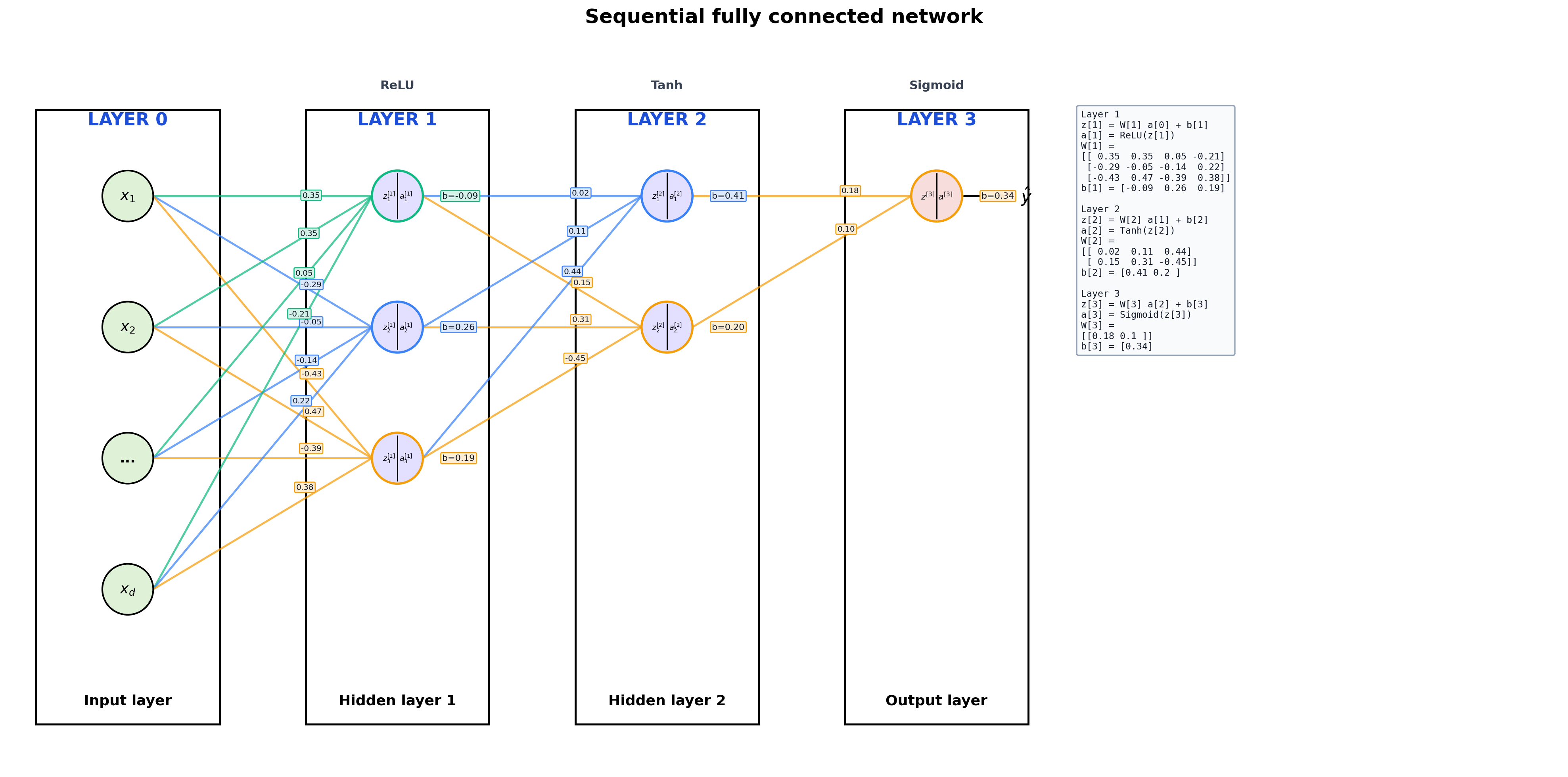
2. A More Typical Sequential Model
The same API works for ordinary lecture examples. It just stops annotating every scalar once the plot would become cluttered.
sequential_model = nn.Sequential(
nn.Linear(4, 3),
nn.ReLU(),
nn.Linear(3, 2),
nn.Tanh(),
nn.Linear(2, 1),
nn.Sigmoid(),
)
print(sequential_model)
print()
print(extract_fully_connected_architecture(sequential_model))
Sequential(
(0): Linear(in_features=4, out_features=3, bias=True)
(1): ReLU()
(2): Linear(in_features=3, out_features=2, bias=True)
(3): Tanh()
(4): Linear(in_features=2, out_features=1, bias=True)
(5): Sigmoid()
)
[LinearLayerSpec(in_features=4, out_features=3, activation='ReLU', name='0'), LinearLayerSpec(in_features=3, out_features=2, activation='Tanh', name='2'), LinearLayerSpec(in_features=2, out_features=1, activation='Sigmoid', name='4')]
fig, ax = visualize_fully_connected_model(
sequential_model,
input_labels=[r"$x_1$", r"$x_2$", "...", r"$x_d$"],
title="Sequential fully connected network",
max_neurons_per_layer=5,
)
plt.show()
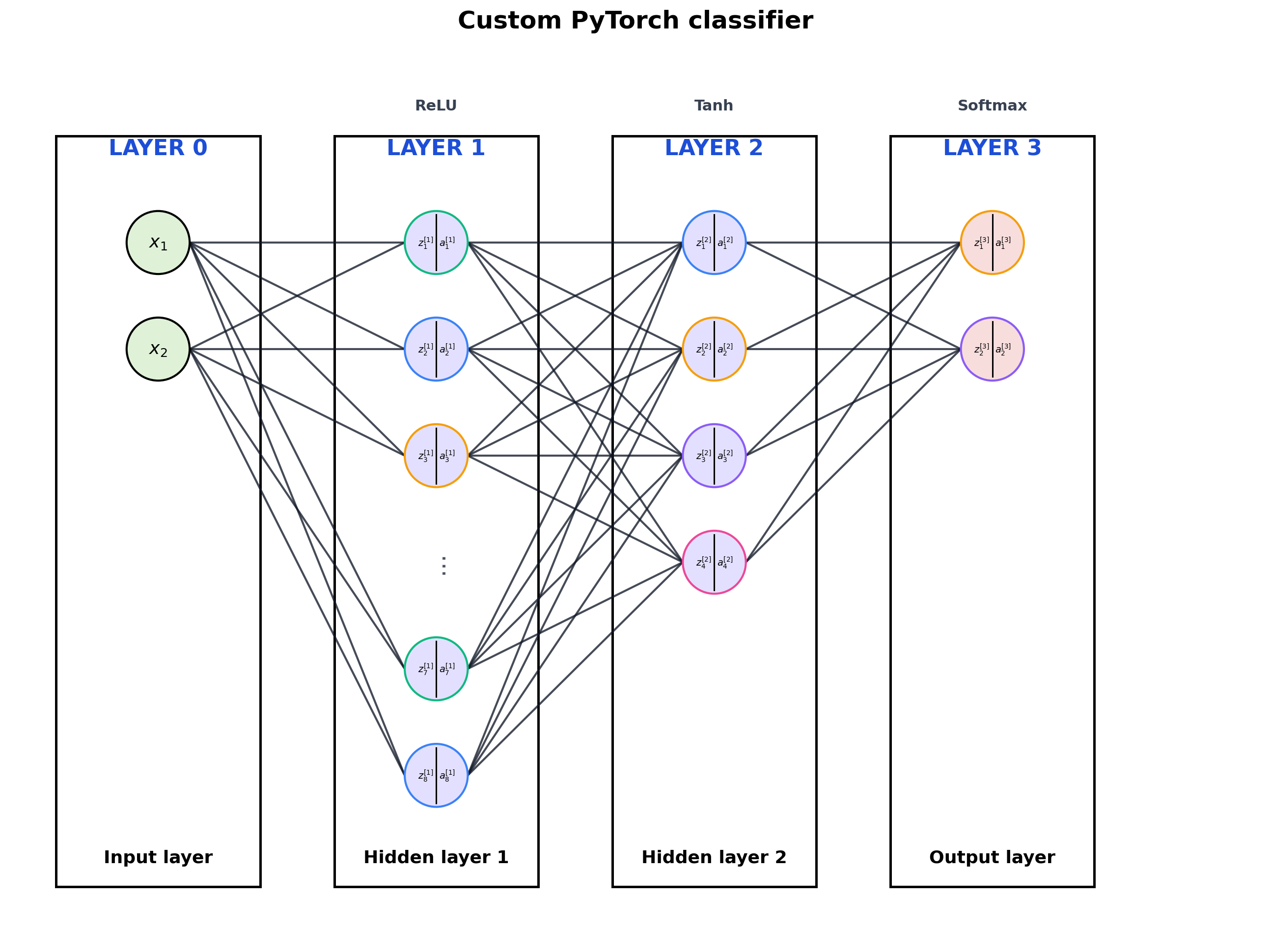
3. Custom nn.Module
The visualizer also works when the network is written as a class with named layers.
class SmallClassifier(nn.Module):
def __init__(self):
super().__init__()
self.hidden1 = nn.Linear(2, 8)
self.hidden2 = nn.Linear(8, 4)
self.output = nn.Linear(4, 2)
def forward(self, x):
x = torch.relu(self.hidden1(x))
x = torch.tanh(self.hidden2(x))
return torch.softmax(self.output(x), dim=-1)
custom_model = SmallClassifier()
print(custom_model)
print()
print(extract_fully_connected_architecture(custom_model))
SmallClassifier(
(hidden1): Linear(in_features=2, out_features=8, bias=True)
(hidden2): Linear(in_features=8, out_features=4, bias=True)
(output): Linear(in_features=4, out_features=2, bias=True)
)
[LinearLayerSpec(in_features=2, out_features=8, activation='ReLU', name='hidden1'), LinearLayerSpec(in_features=8, out_features=4, activation='Tanh', name='hidden2'), LinearLayerSpec(in_features=4, out_features=2, activation='Softmax', name='output')]
fig, ax = visualize_fully_connected_model(
custom_model,
input_labels=[r"$x_1$", r"$x_2$"],
output_labels=["class 1", "class 2"],
title="Custom PyTorch classifier",
max_neurons_per_layer=6,
)
plt.show()
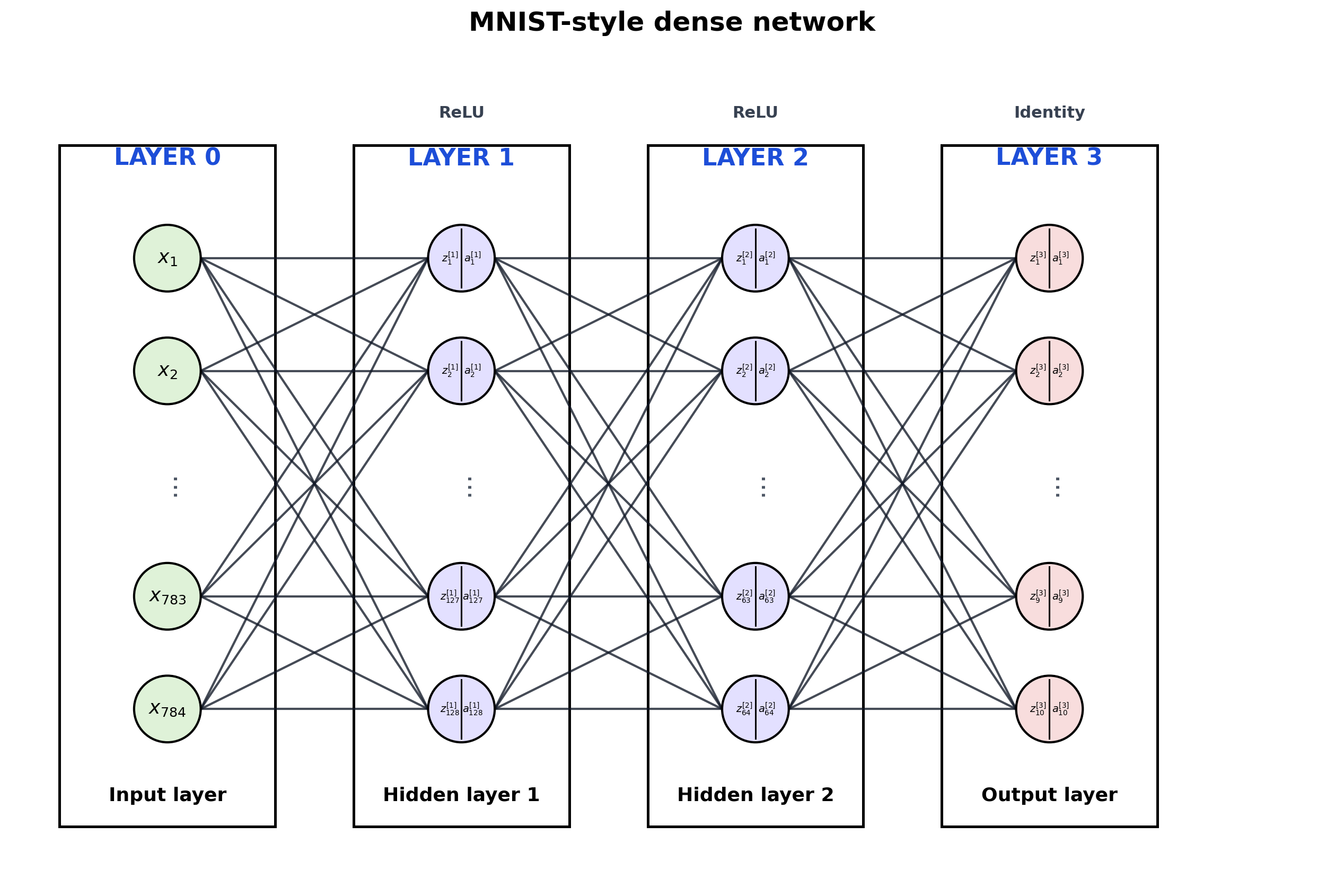
4. Wide Model with Truncation
Large dense layers get summarized with ... so the diagram stays readable.
wide_model = nn.Sequential(
nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
)
fig, ax = visualize_fully_connected_model(
wide_model,
title="MNIST-style dense network",
max_neurons_per_layer=5,
)
plt.show()
5. Save a Diagram
Reuse the figure in slides or notes by saving PNG and SVG.
output_dir = repo_root / "outputs"
png_path = output_dir / "fc_visualizer_demo.png"
svg_path = output_dir / "fc_visualizer_demo.svg"
fig, ax = visualize_fully_connected_model(
small_model,
input_labels=[r"$x_1$", r"$x_2$"],
input_vector=x_example,
title="Saved demo network",
show_edge_weights=True,
show_biases=True,
show_values=True,
show_matrix_details=True,
)
save_figure(fig, png_path=png_path, svg_path=svg_path)
print(f"Saved PNG: {png_path}")
print(f"Saved SVG: {svg_path}")
Saved PNG: /Users/nipun/git/principles-ai-teaching/outputs/fc_visualizer_demo.png
Saved SVG: /Users/nipun/git/principles-ai-teaching/outputs/fc_visualizer_demo.svg
Notes
The visualizer traces the forward graph and looks for:
nn.Linear- activations such as
ReLU, Tanh, Sigmoid, Softmax
- simple reshaping or flattening operations
Small-network extras:
input_vector=... computes and annotates forward-pass valuesshow_edge_weights=True writes each scalar weight on its edgeshow_biases=True shows each bias near its neuronshow_matrix_details=True adds a side panel with W, b, z, and a
If the model contains convolution, skip connections, attention, concatenation, or branching, the visualizer will reject it on purpose.