Code
import autograd.numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import GPyNipun Batra
September 3, 2021
In our previous post we had mentioned (for the noiseless case):
Given train data \[ D=\left(x_{i}, y_{i}\right), i=1: N \] Given a test set \(X_{*}\) of size \(N_{*} \times d\) containing \(N_{*}\) points in \(\mathbb{R}^{d},\) we want to predict function outputs \(y_{*}\) We can write: \[ \left(\begin{array}{l} y \\ y_{*} \end{array}\right) \sim \mathcal{N}\left(\left(\begin{array}{l} \mu \\ \mu_{*} \end{array}\right),\left(\begin{array}{cc} K & K_{*} \\ K_{*}^{T} & K_{* *} \end{array}\right)\right) \] where \[ \begin{aligned} K &=\operatorname{Ker}(X, X) \in \mathbb{R}^{N \times N} \\ K_{*} &=\operatorname{Ker}\left(X, X_{*}\right) \in \mathbb{R}^{N \times N} \\ K_{* *} &=\operatorname{Ker}\left(X_{*}, X_{*}\right) \in \mathbb{R}^{N_{*} \times N_{*}} \end{aligned} \]
Thus, from the property of conditioning of multivariate Gaussian, we know that:
\[y \sim \mathcal{N}_N(\mu, K)\]
We will assume \(\mu\) to be zero. Thus, we have for the train data, the following expression:
\[y \sim \mathcal{N}_N(0, K)\]
For the noisy case, we have:
\[y \sim \mathcal{N}_N(0, K + \sigma_{noise}^2\mathcal{I}_N)\]
From this expression, we can write the log-likelihood of data computed over the kernel parameters \(\theta\) as:
\[\mathcal{LL}(\theta) = \log(\frac{\exp((-1/2)(y-0)^T (K+\sigma_{noise}^2\mathcal{I}_N)^{-1}(y-0))}{(2\pi)^{N/2}|(K+\sigma_{noise}^2\mathcal{I}_N)|^{1/2}})\]
Thus, we can write:
\[\mathcal{LL}(\theta) =\log P(\mathbf{y} | X, \theta)=-\frac{1}{2} \mathbf{y}^{\top} M^{-1} \mathbf{y}-\frac{1}{2} \log |M|-\frac{N}{2} \log 2 \pi\]
where \[M = K + \sigma_{noise}^2\mathcal{I}_N\]
As before, we will be using the excellent Autograd library for automatically computing the gradient of an objective function with respect to the parameters. We will also be using GPy for verifying our calculations.
Let us start with some basic imports.
The definition of the (1-dimensional) RBF kernel has a Gaussian-form, defined as:
\[ \kappa_\mathrm{rbf}(x_1,x_2) = \sigma^2\exp\left(-\frac{(x_1-x_2)^2}{2\mathscr{l}^2}\right) \]
Looks good. Our function is matching GPy’s kernel.
# lambda function, call f(x) to generate data
f = lambda x: 0.4*x**2 - 0.15*x**3 + 0.5*x**2 - 0.002*x**5 + 0.0002*x**6 +0.5*(x-2)**2
n = 50
np.random.seed(0)
X = np.linspace(0.05, 4.95, n)[:,None]
Y = f(X) + np.random.normal(0., 0.1, (n,1)) # note that np.random.normal takes mean and s.d. (not variance), 0.1^2 = 0.01
plt.plot(X, Y, "kx", mew=0.5, label='Train points')
plt.xlabel("x"), plt.ylabel("f")
plt.legend();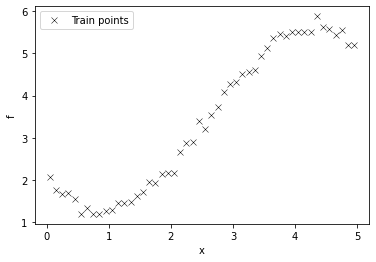
Based on our above mentioned theory, we can now write the NLL function as follows
We will now compare the NLL from our method with GPy for a fixed set of parameters
Name : GP regression Objective : 70.98928950981251 Number of Parameters : 3 Number of Optimization Parameters : 3 Updates : True Parameters: GP_regression. | value | constraints | priors rbf.variance | 1.0 | +ve | rbf.lengthscale | 1.0 | +ve | Gaussian_noise.variance | 1.0 | +ve |
Excellent, we can see that our method gives the same NLL. Looks like we are on the right track! One caveat here is that I have set the normalizer to be False, which means that GPy will not be mean centering the data.
We will now use GPy to optimize the GP parameters
Name : GP regression Objective : -20.145595097323056 Number of Parameters : 3 Number of Optimization Parameters : 3 Updates : True Parameters: GP_regression. | value | constraints | priors rbf.variance | 38.67759006151069 | +ve | rbf.lengthscale | 2.9466566795409825 | +ve | Gaussian_noise.variance | 0.012471600151185298 | +ve |
It seems that variance close to 28 and length scale close to 2.7 give the optimum objective for the GP
We will now plot the NLL obtained from our calculations as a function of variance and lengthscale. For comparing our solution with GPy solution, I will be setting noise variance to be 0.0075
We will now try to find the “optimum” \(\sigma\) and lengthscale from this NLL space.
batch_size = 20
idx = np.random.randint(X.shape[0], size=batch_size)
X_subset, Y_subset = X[idx, :], Y[idx]
X_subset, Y_subset
plt.plot(X, Y, "kx", mew=2, label='Train points')
plt.plot(X_subset, Y_subset, "go", mew=2, label='Batch of Train points')
plt.legend()
plt.title("NLL = %0.3f" %nll(X_subset, Y_subset, 1, 1, 1))Text(0.5, 1.0, 'NLL = 40.707')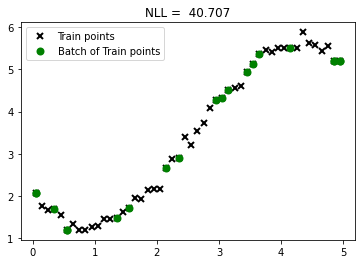
sigma = 2.
l = 2.
noise = 0.5
lr = 1e-2
num_iter = 1500
np.random.seed(0)
nll_arr = np.zeros(num_iter)
for iteration in range(num_iter):
idx = np.random.randint(X.shape[0], size=batch_size)
X_subset, Y_subset = X[idx, :], Y[idx]
nll_arr[iteration] = nll(X_subset, Y_subset, sigma, l, noise)
del_sigma, del_l, del_noise = grad_objective(X_subset, Y_subset, sigma, l, noise)
sigma = sigma - lr*del_sigma
l = l - lr*del_l
noise = noise - lr*del_noise
lr = lr/(iteration+1)sigma = 2.
l = 2.
noise = 0.5
lr = 1e-2
num_iter = 1500
np.random.seed(0)
nll_arr = np.zeros(num_iter)
for iteration in range(num_iter):
# Sample 1 point
idx = np.random.randint(X.shape[0], size=1)
x_val = X[idx]
K = batch_size
a = np.abs(X - x_val).flatten()
idx = np.argpartition(a,K)[:K]
X_subset = X[idx, :]
Y_subset = Y[idx, :]
nll_arr[iteration] = nll(X_subset, Y_subset, sigma, l, noise)
del_sigma, del_l, del_noise = grad_objective(X_subset, Y_subset, sigma, l, noise)
sigma = sigma - lr*del_sigma
l = l - lr*del_l
noise = noise - lr*del_noise
lr = lr/(iteration+1)sigma = 2.
l = 2.
noise = 0.5
lr = 1e-2
num_iter = 100
batch_size = 5
nll_arr = np.zeros(num_iter)
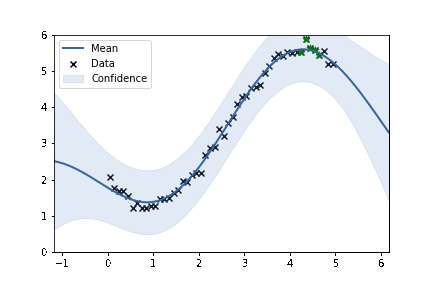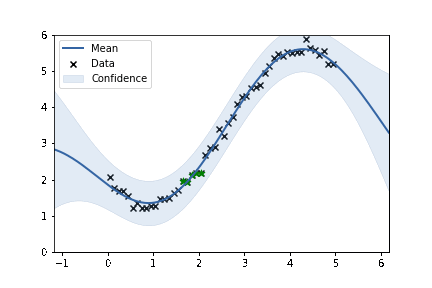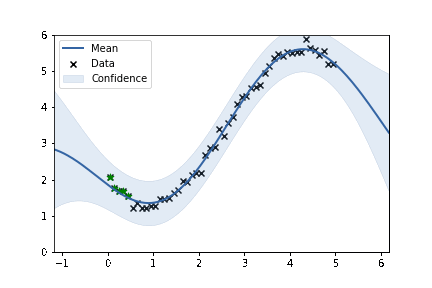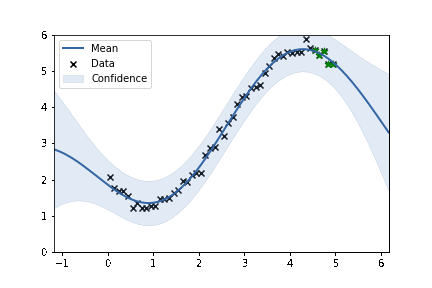
fig, ax = plt.subplots()
np.random.seed(0)
for iteration in range(num_iter):
# Sample 1 point
idx = np.random.randint(X.shape[0], size=1)
x_val = X[idx]
K = batch_size
a = np.abs(X - x_val).flatten()
idx = np.argpartition(a,K)[:K]
X_subset = X[idx, :]
Y_subset = Y[idx, :]
nll_arr[iteration] = nll(X_subset, Y_subset, sigma, l, noise)
del_sigma, del_l, del_noise = grad_objective(X_subset, Y_subset, sigma, l, noise)
sigma = sigma - lr*del_sigma
l = l - lr*del_l
noise = noise - lr*del_noise
k.lengthscale = l
k.variance = sigma**2
m = GPy.models.GPRegression(X, Y, k, normalizer=False)
m.Gaussian_noise = noise**2
m.plot(ax=ax, alpha=0.2)['dataplot'];
ax.scatter(X_subset, Y_subset, color='green', marker='*', s = 50)
plt.ylim((0, 6))
#plt.title(f"Iteration: {iteration:04}, Objective :{nll_arr[iteration]:04.2f}")
plt.savefig(f"/Users/nipun/Desktop/gp_learning/{iteration:04}.png")
ax.clear()
lr = lr/(iteration+1)
plt.clf()<Figure size 432x288 with 0 Axes>