Code
import numpy as npimport matplotlib.pyplot as pltimport tensorflow as tfimport seaborn as snsimport tensorflow_probability as tfpimport pandas as pd= tfp.distributions= 'talk' ,font_scale= 1 )% matplotlib inline% config InlineBackend.figure_format= 'retina'
Univariate normal
Code
= tfd.Normal(loc= 0. , scale= 1. )
Code
<tfp.distributions.Normal 'Normal' batch_shape=[] event_shape=[] dtype=float32>
Code
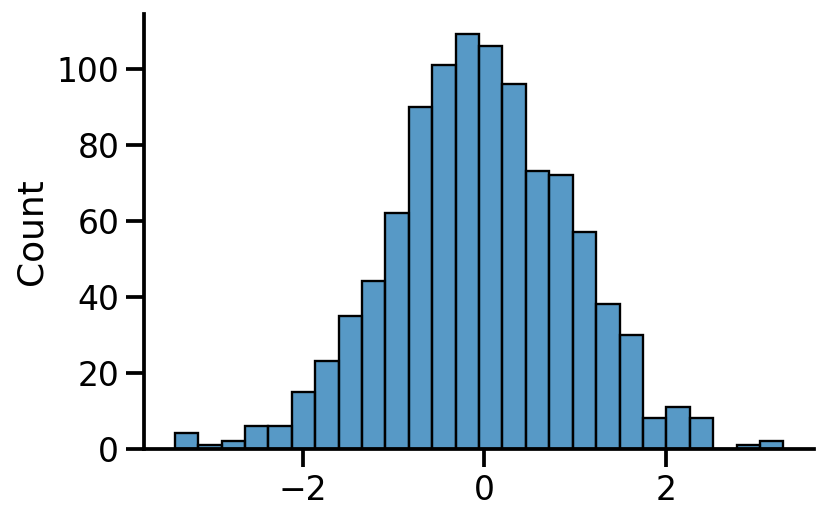
= uv_normal.sample(1000 )
Code
Code
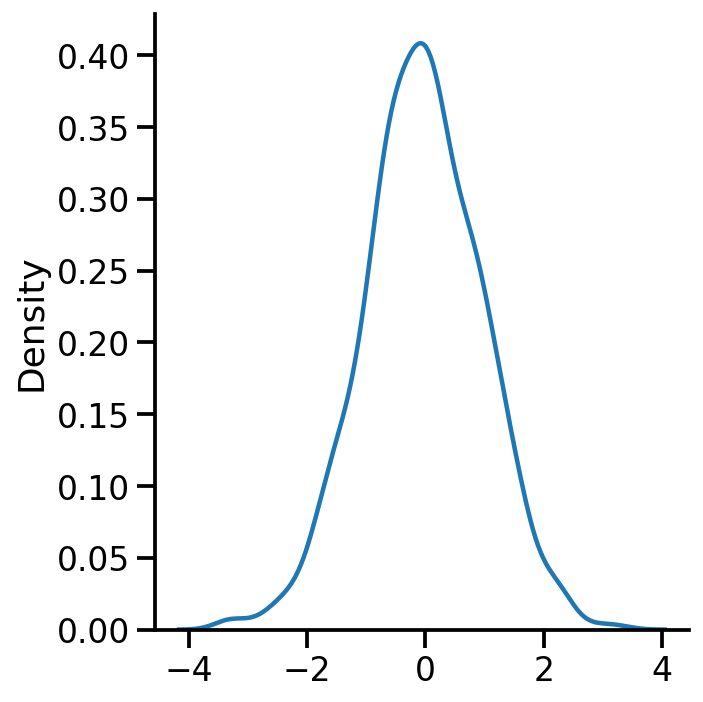
= 'kde' )
Code
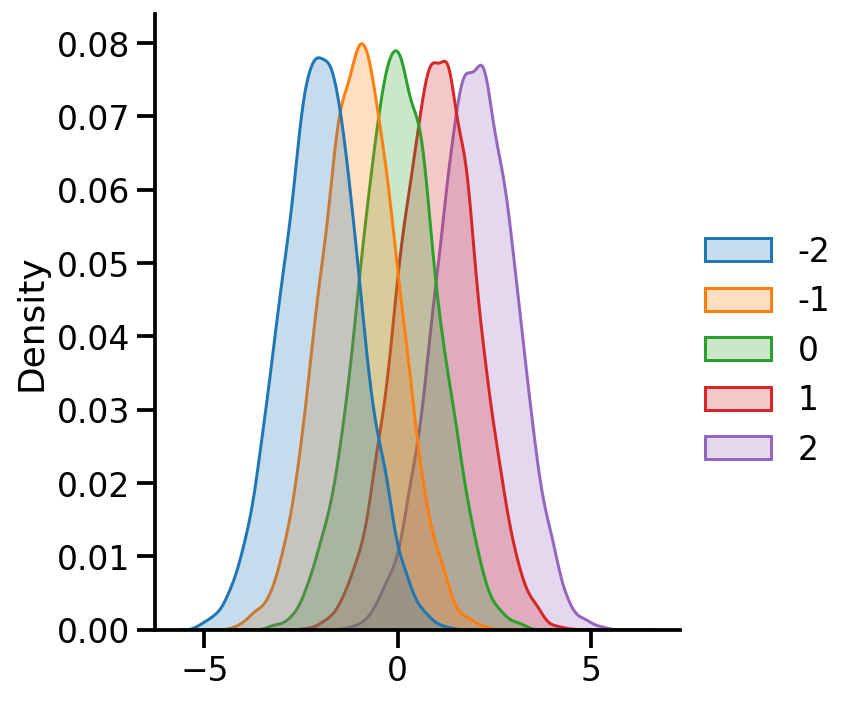
= {x: tfd.Normal(loc= x, scale= 1. ) for x in [- 2 , - 1 , 0 , 1 , 2 ]}
Code
= pd.DataFrame({x:uv_normal_dict_mean[x].sample(10000 ).numpy() for x in uv_normal_dict_mean})
Code
= 'kde' , fill= True )
Code
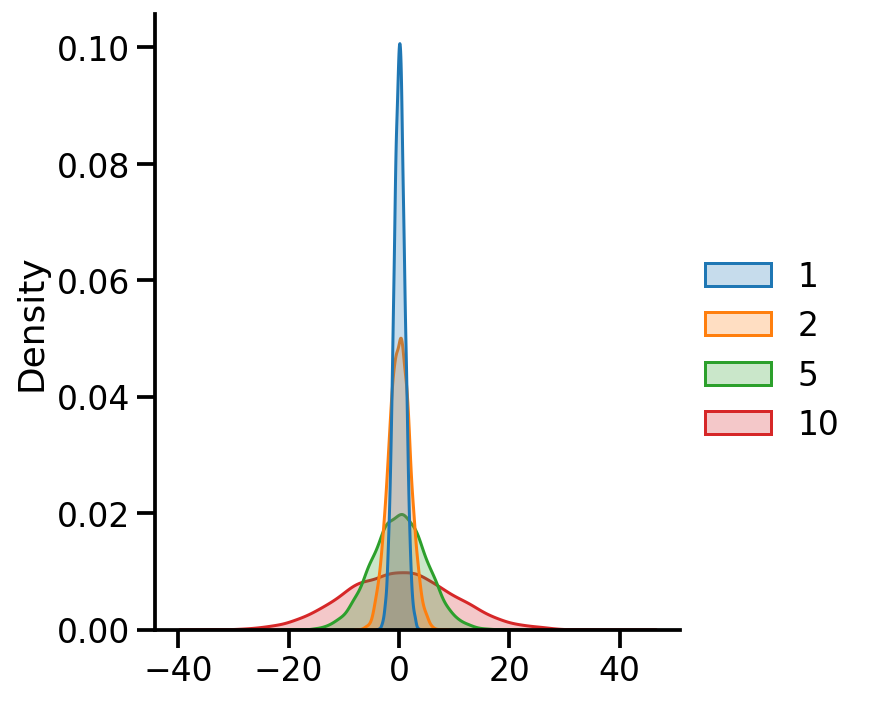
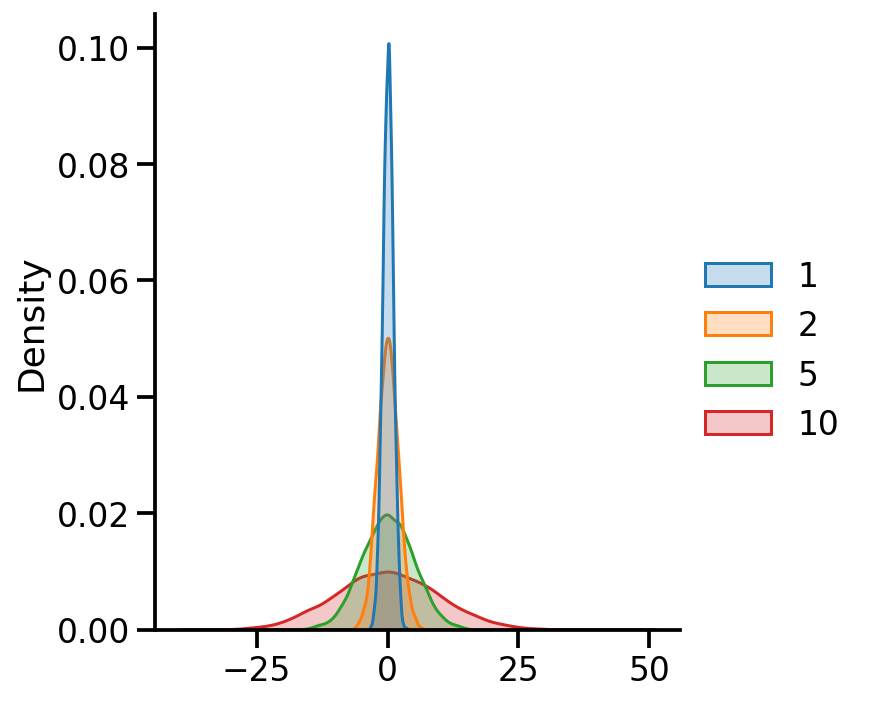
= {x: tfd.Normal(loc= 0 , scale= x) for x in [1 , 2 , 5 , 10 ]}= pd.DataFrame({x:uv_normal_dict_var[x].sample(10000 ).numpy() for x in uv_normal_dict_var})
Code
= 'kde' , fill= True )
Using batches
Code
= pd.DataFrame(= [0. , 0. , 0. , 0. ],= [1. , 2. , 5. , 10. ]).sample(10000 ).numpy())= [1 , 2 , 5 , 10 ]= 'kde' , fill= True )
Code
= [0. , 0. , 0. , 0. ],= [1. , 2. , 5. , 10. ])
<tfp.distributions.Normal 'Normal' batch_shape=[4] event_shape=[] dtype=float32>
Code
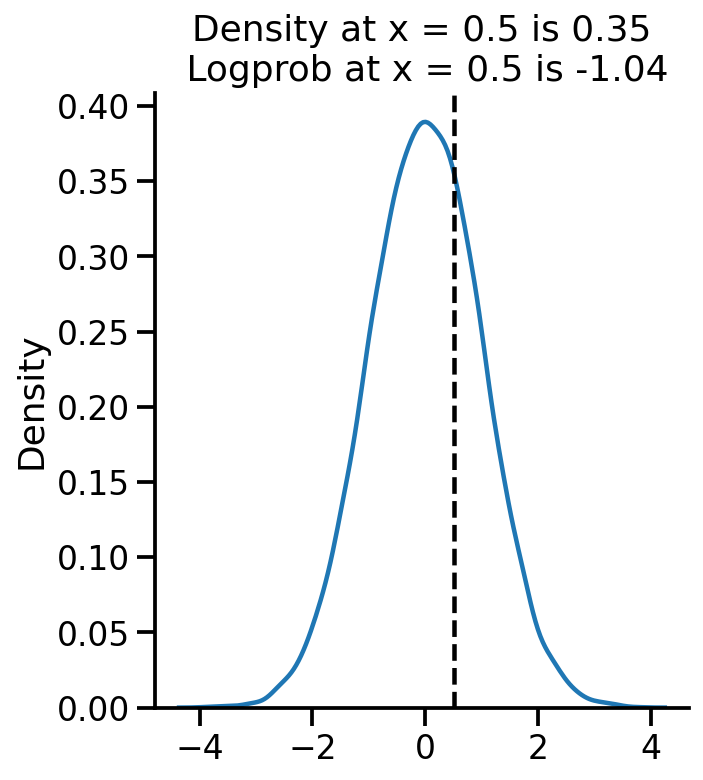
= uv_normal.sample(10000 )= 'kde' )0.5 , color= 'k' , linestyle= '--' )= uv_normal.prob(0.5 ).numpy()= uv_normal.log_prob(0.5 ).numpy()"Density at x = 0.5 is {:.2f} \n Logprob at x = 0.5 is {:.2f} " .format (pdf_05, log_pdf_05))
Text(0.5, 1.0, 'Density at x = 0.5 is 0.35\n Logprob at x = 0.5 is -1.04')
Learning parameters
Let us generate some normally distributed data and see if we can learn the mean.
Code
= uv_normal.sample(10000 )
Code
(<tf.Tensor: shape=(), dtype=float32, numpy=0.0>,
<tf.Tensor: shape=(), dtype=float32, numpy=1.0>)
Let us create a new TFP trainable distribution where we wish to learn the mean.
Code
= tfd.Normal(loc = tf.Variable(- 1. , name= 'loc' ), scale = 1. )
Code
<tfp.distributions.Normal 'Normal' batch_shape=[] event_shape=[] dtype=float32>
Code
(<tf.Variable 'loc:0' shape=() dtype=float32, numpy=-1.0>,)
Code
(<tf.Tensor: shape=(), dtype=float32, numpy=-0.024403999>,
<tf.Tensor: shape=(), dtype=float32, numpy=0.9995617>)
Code
def nll(train):return - tf.reduce_mean(to_train.log_prob(train))
Code
<tf.Tensor: shape=(), dtype=float32, numpy=1.8946133>
Code
def get_loss_and_grads(train):with tf.GradientTape() as tape:= nll(train)= tape.gradient(loss, to_train.trainable_variables)return loss, grads
Code
(<tf.Tensor: shape=(), dtype=float32, numpy=1.8946133>,
(<tf.Tensor: shape=(), dtype=float32, numpy=-0.97559595>,))
Code
= tf.keras.optimizers.Adam(learning_rate= 0.01 )
Code
<keras.optimizer_v2.adam.Adam at 0x7f94c97ae490>
Code
= 500 = np.empty(iterations)= np.empty(iterations)for i in range (iterations):= get_loss_and_grads(train_data)= loss= to_train.trainable_variables[0 ].numpy()zip (grads, to_train.trainable_variables))if i% 50 == 0 :print (i, loss.numpy())
0 1.8946133
50 1.5505791
100 1.4401271
150 1.4205703
200 1.4187955
250 1.4187206
300 1.4187194
350 1.4187193
400 1.4187193
450 1.4187194
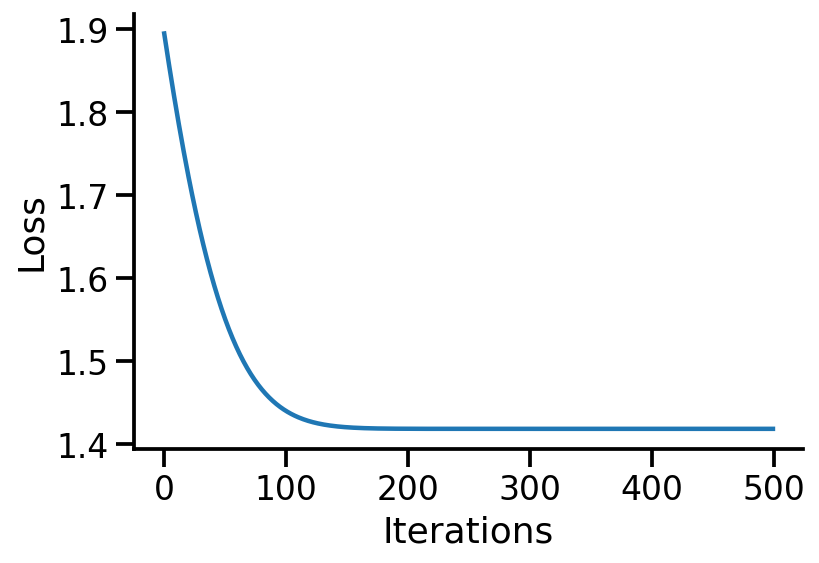
Code
"Iterations" )"Loss" )
Code
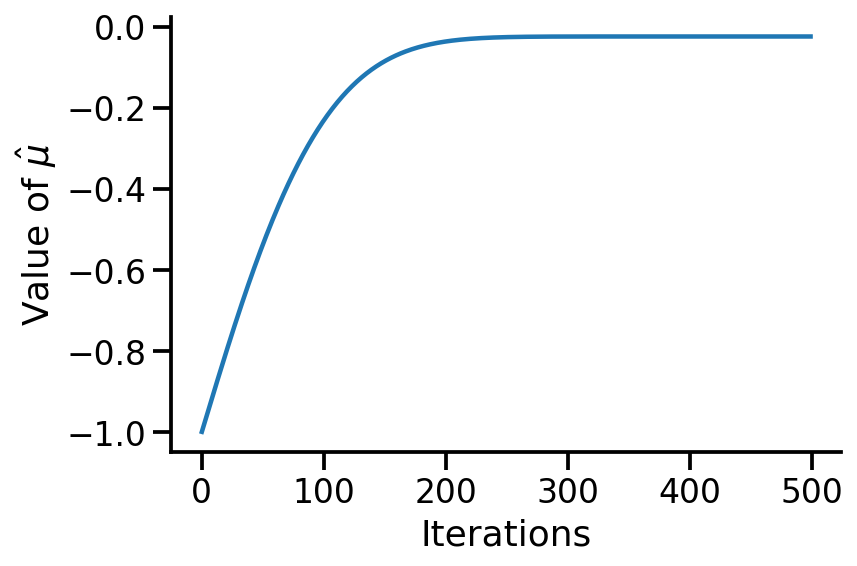
"Iterations" )r"Value of $ \ hat{ \m u} $ " )
Text(0, 0.5, 'Value of $\\hat{\\mu}$')
Code
= tfd.Normal(loc = tf.Variable(- 1. , name= 'loc' ), scale = tf.Variable(10. , name= 'scale' ))def nll(train):return - tf.reduce_mean(to_train_mean_var.log_prob(train))def get_loss_and_grads(train):with tf.GradientTape() as tape:= nll(train)= tape.gradient(loss, to_train_mean_var.trainable_variables)return loss, grads= tf.keras.optimizers.Adam(learning_rate= 0.01 )= 1000 = np.empty(iterations)= np.empty(iterations)= np.empty(iterations)for i in range (iterations):= get_loss_and_grads(train_data)= loss= to_train_mean_var.trainable_variables[0 ].numpy()= to_train_mean_var.trainable_variables[1 ].numpy()zip (grads, to_train_mean_var.trainable_variables))if i% 50 == 0 :print (i, loss.numpy())
0 3.2312806
50 3.1768403
100 3.1204312
150 3.0602157
200 2.9945102
250 2.9219644
300 2.8410006
350 2.749461
400 2.6442661
450 2.5208094
500 2.3718355
550 2.1852348
600 1.9403238
650 1.6161448
700 1.4188237
750 1.4187355
800 1.4187193
850 1.4187193
900 1.4187193
950 1.4187193
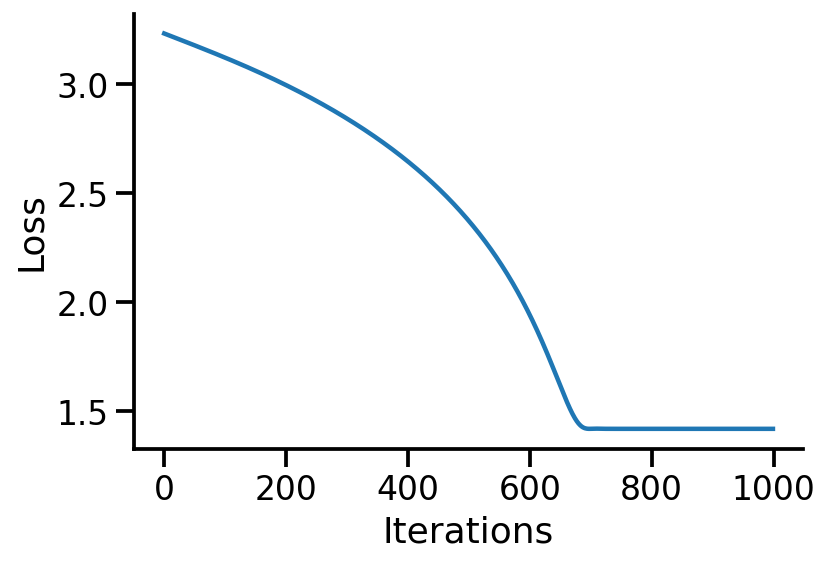
Code
"Iterations" )"Loss" )
Code
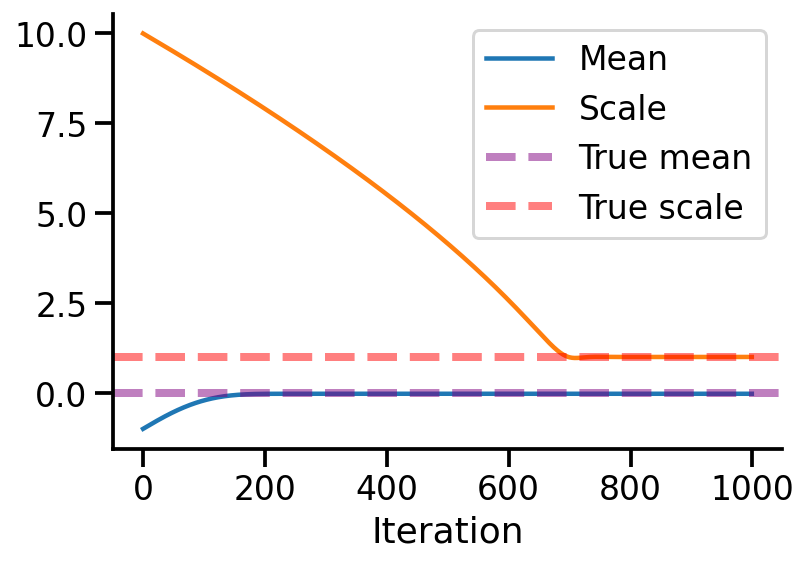
= pd.DataFrame({"Mean" :vals_means, "Scale" :vals_scale}, index= range (iterations))= 'Iteration'
Code
= 1 )0 , linestyle= '--' , lw = 4 , label = 'True mean' , alpha= 0.5 , color= 'purple' )1 , linestyle= '--' , lw = 4 , label = 'True scale' , alpha= 0.5 , color= 'red' )
Multivariate Normal
Code
= tfd.MultivariateNormalFullCovariance(loc= [0 , 0 ], covariance_matrix= [[1 , 0.5 ], [0.5 , 2 ]])
Code
= pd.DataFrame(mv_normal.sample(10000 ).numpy())= [r' $ x_1 $ ' , r' $ x_2 $ ' ]
Code
<tf.Tensor: shape=(), dtype=float32, numpy=0.120309845>
Code
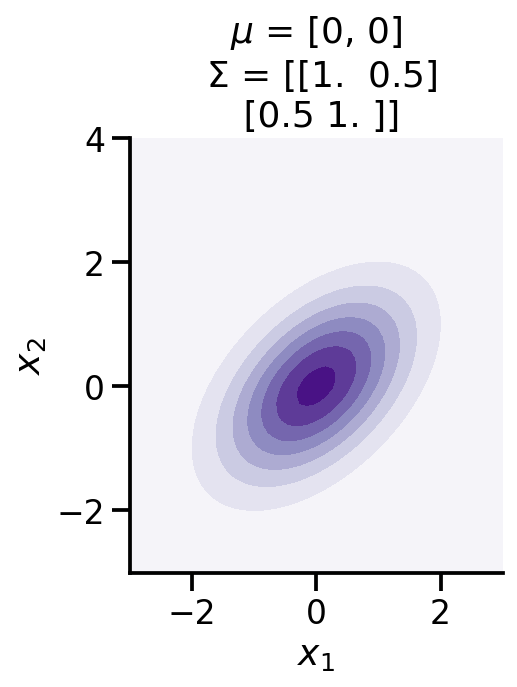
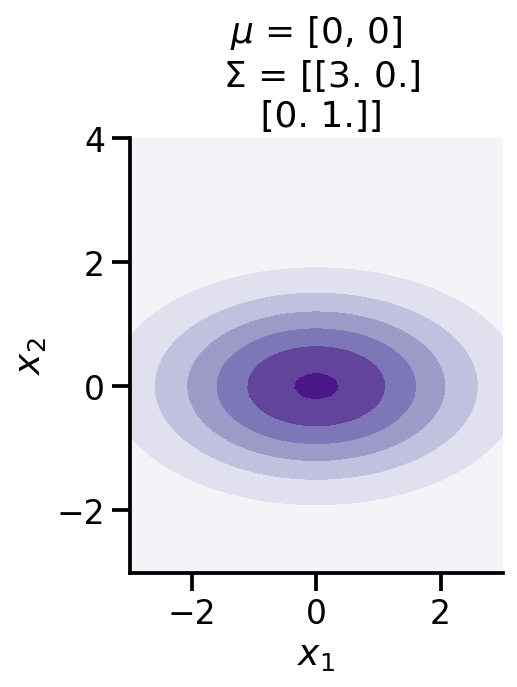
from mpl_toolkits.mplot3d import Axes3Dfrom matplotlib import cmdef make_pdf_2d_gaussian(mu, sigma):= 60 = np.linspace(- 3 , 3 , N)= np.linspace(- 3 , 4 , N)= np.meshgrid(X, Y)# Pack X and Y into a single 3-dimensional array = np.empty(X.shape + (2 ,))0 ] = X1 ] = Y= tfd.MultivariateNormalFullCovariance(loc= mu, covariance_matrix= sigma)= F.prob(pos)= cm.Purples)r" $ x_1 $ " )r" $ x_2 $ " )'equal' )f'$ \ mu$ = { mu} \n $ \ Sigma$ = { np. array(sigma)} ' )
Code
0 , 0 ,], [[1 , 0.5 ,], [0.5 , 1 ]])
Code
0 , 0 ,], [[3 , 0. ,], [0. , 1 ]])
Code
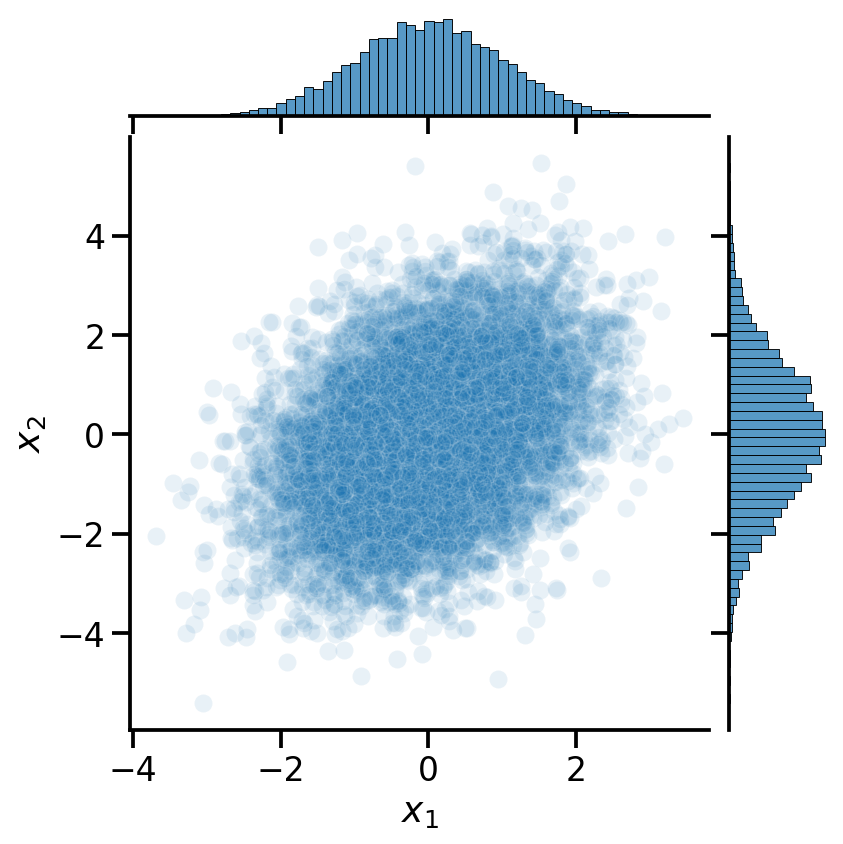
= mv_data,= r' $ x_1 $ ' ,y= r' $ x_2 $ ' ,= 0.1 )
Code
0
2.155621
-0.343866
1
-0.731184
0.378393
2
0.832593
-0.459740
3
-0.701200
-0.249675
4
-0.430790
-1.694002
...
...
...
9995
-0.165910
-0.171243
9996
0.208389
-1.698432
9997
-0.030418
0.353905
9998
1.342328
1.127457
9999
-0.145741
0.830713
10000 rows × 2 columns