Code
import numpy as npimport sklearnimport matplotlib.pyplot as pltimport pandas as pd% matplotlib inline
https://towardsdatascience.com/introduction-to-reliability-diagrams-for-probability-calibration-ed785b3f5d44
Code
= np.array([0.9 , 0.2 , 0.7 , 0.4 , 0.8 , 0.1 , 0.2 , 0.8 , 0.5 , 0.9 ])
Code
= np.ones_like(p)1 , 6 , 7 , 8 ]] = 0
array([1., 0., 1., 1., 1., 1., 0., 0., 0., 1.])
Code
= np.linspace(0 , 1 , num_splits + 1 )= [(x, y) for (x, y) in zip (splits_arr[:- 1 ], splits_arr[1 :])]
[(0.0, 0.3333333333333333),
(0.3333333333333333, 0.6666666666666666),
(0.6666666666666666, 1.0)]
Code
= splits_arr)
0 (0.667, 1.0]
1 (0.0, 0.333]
2 (0.667, 1.0]
3 (0.333, 0.667]
4 (0.667, 1.0]
5 (0.0, 0.333]
6 (0.0, 0.333]
7 (0.667, 1.0]
8 (0.333, 0.667]
9 (0.667, 1.0]
dtype: category
Categories (3, interval[float64, right]): [(0.0, 0.333] < (0.333, 0.667] < (0.667, 1.0]]
Code
= np.digitize(p, splits_arr)
array([3, 1, 3, 2, 3, 1, 1, 3, 2, 3])
Code
= {}= {}for group in np.unique(splits):= p[splits== group]#frac_pos[group] = true_labels[splits==group].sum()*1.0/len(p_group[group]) = true_labels[splits== group]#print(np.arange(10)[splits==group])
Code
{1: array([0.2, 0.1, 0.2]),
2: array([0.4, 0.5]),
3: array([0.9, 0.7, 0.8, 0.8, 0.9])}
Code
{1: array([0., 1., 0.]), 2: array([1., 0.]), 3: array([1., 1., 1., 0., 1.])}
Code
= {k:np.mean(v) for k, v in p_group.items()}
{1: 0.16666666666666666, 2: 0.45, 3: 0.8200000000000001}
Code
= {k:np.sum (v)* 1.0 / len (v) for k, v in labels_pos.items()}
{1: 0.3333333333333333, 2: 0.5, 3: 0.8}
Code
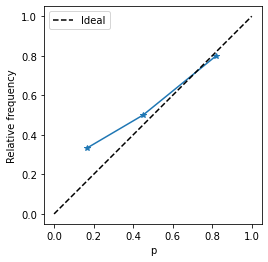
= '*' )- 0.05 , 1.05 ))- 0.05 , 1.05 ))"equal" )"p" )"Relative frequency" )0 , 1 ], [0 , 1 ], color= 'k' , ls= '--' , label= 'Ideal' )
Let us wrap into a function
Code
def calib_curve(true, pred, n_bins = 10 ):= np.linspace(0 , 1 , n_bins + 1 )= np.digitize(pred, splits_arr)= {}= {}for group in np.unique(splits):= pred[splits== group]= true[splits== group]= {k:np.mean(v) for k, v in p_group.items()}= {k:np.sum (v)* 1.0 / len (v) for k, v in labels_pos.items()}= np.array([len (v) for v in labels_pos.values()])return np.array(list (p_group_mean.values())), np.array(list (fracs.values())), counts
Code
from sklearn.calibration import calibration_curve, CalibrationDisplay= calibration_curve(true_labels, p, n_bins= 3 )
Code
array([0.33333333, 0.5 , 0.8 ])
Code
array([0.16666667, 0.45 , 0.82 ])
Code
= calib_curve(true_labels, p, 3 )
array([0.16666667, 0.45 , 0.82 ])
Expected Calibration Error
Code
abs (p_ours- p_hat_ours)* count).mean()
Code
from sklearn.datasets import make_classification
Code
= make_classification(n_features= 2 , n_informative= 2 , n_redundant= 0 , random_state= 0 )
Code
0 ], X[:, 1 ], c = y)
Code
from sklearn.linear_model import LogisticRegression
Code
= LogisticRegression()
Code
LogisticRegression() In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
Code
= CalibrationDisplay.from_estimator(= 11 ,
Code
= lr.predict_proba(X)[:, 1 ]
Code
= calib_curve(y, pred_p, 11 )
Code
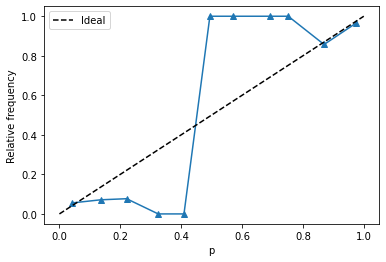
= '^' )"p" )"Relative frequency" )0 , 1 ], [0 , 1 ], color= 'k' , ls= '--' , label= 'Ideal' )
Code
abs (probs- fractions)* counts).mean()
Code
array([18, 14, 13, 4, 2, 4, 2, 2, 6, 7, 28])