SUPPORTED_AGGREGATES = {"mean", "sum", "count", "median", "min", "max"}
def _mark_type(spec):
mark = spec.get("mark")
return mark if isinstance(mark, str) else mark.get("type")
def _enc_label(enc_channel):
if "title" in enc_channel:
return enc_channel["title"]
if "aggregate" in enc_channel and enc_channel.get("field"):
return f"{enc_channel['aggregate']}({enc_channel['field']})"
return enc_channel.get("field", "")
def _has_facet(spec):
enc = spec.get("encoding", {})
return "column" in enc or "row" in enc or "facet" in spec
class VegaLiteToMatplotlib:
"""Recursive Vega-Lite -> Matplotlib code translator.
Dispatch:
top-level -> visit_layer | visit_facet | visit_unit
mark -> visit_mark_bar | visit_mark_point | visit_mark_circle
| visit_mark_line | visit_mark_area | visit_mark_tick
"""
def __init__(self, spec):
self.spec = spec
self._lines = []
self._indent = 0
# --- emit / indent ---
def emit(self, *lines):
for line in lines:
self._lines.append((" " * self._indent + line) if line else "")
def push_indent(self):
self._indent += 1
def pop_indent(self):
self._indent -= 1
# --- entry point ---
def code(self):
if self._lines:
return "\n".join(self._lines)
self.emit("import matplotlib.pyplot as plt")
self.emit("import numpy as np")
self.emit("import pandas as pd")
self.emit("")
spec = self.spec
if _has_facet(spec):
self.visit_facet(spec)
elif "layer" in spec:
self.emit("fig, ax = plt.subplots(figsize=(6, 3.6))")
self.emit("")
self.visit_layer(spec, ax_expr="ax")
self.emit("fig.tight_layout()")
else:
self.emit("fig, ax = plt.subplots(figsize=(6, 3.6))")
self.emit("")
self.visit_unit(spec, ax_expr="ax")
self.emit("fig.tight_layout()")
self.emit("plt.show()")
return "\n".join(self._lines)
# --- top-level visitors ---
def visit_unit(self, spec, ax_expr, df_expr=None):
if df_expr is None:
df_expr = "df"
self._emit_data(spec, df_expr)
self._emit_transforms(spec, df_expr)
self._dispatch_mark(spec, ax_expr, df_expr)
self._emit_axes(spec, ax_expr)
def visit_layer(self, spec, ax_expr):
outer_data = spec.get("data")
for i, layer in enumerate(spec["layer"]):
df_var = f"df_layer_{i}"
inner = dict(layer)
inner.setdefault("data", outer_data)
self._emit_data(inner, df_var)
self._emit_transforms(inner, df_var)
self._dispatch_mark(inner, ax_expr, df_var)
# axes from the first layer
self._emit_axes(spec["layer"][0], ax_expr)
if spec.get("title"):
self.emit(f"{ax_expr}.set_title({_unwrap_title(spec['title'])!r})")
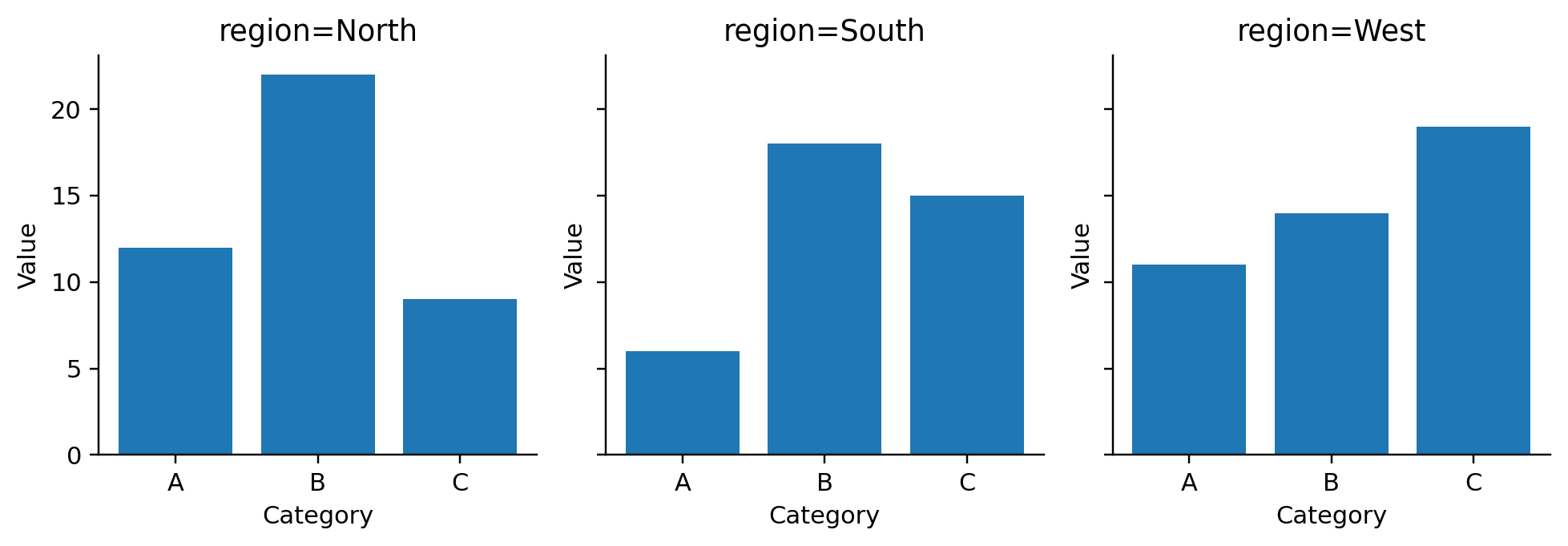
def visit_facet(self, spec):
enc = spec.get("encoding", {})
col_field = enc.get("column", {}).get("field")
row_field = enc.get("row", {}).get("field")
if col_field is None and row_field is None:
raise NotImplementedError("Facet requires column or row encoding")
if col_field and row_field:
raise NotImplementedError("2D faceting (column AND row) not implemented")
self._emit_data(spec, "df")
self._emit_transforms(spec, "df")
facet_field = col_field or row_field
is_column = col_field is not None
self.emit(f"facet_values = list(dict.fromkeys(df[{facet_field!r}]))")
if is_column:
self.emit("fig, axes = plt.subplots(")
self.emit(" 1, len(facet_values),")
self.emit(" figsize=(3.0 * len(facet_values), 3.2),")
self.emit(" sharey=True,")
self.emit(")")
else:
self.emit("fig, axes = plt.subplots(")
self.emit(" len(facet_values), 1,")
self.emit(" figsize=(5.0, 2.6 * len(facet_values)),")
self.emit(" sharex=True,")
self.emit(")")
self.emit("if len(facet_values) == 1:")
self.push_indent(); self.emit("axes = [axes]"); self.pop_indent()
self.emit("for ax, facet_val in zip(axes, facet_values):")
self.push_indent()
self.emit(f"sub = df[df[{facet_field!r}] == facet_val]")
inner = dict(spec)
inner_enc = dict(enc)
inner_enc.pop("column", None)
inner_enc.pop("row", None)
inner["encoding"] = inner_enc
inner.pop("data", None)
self.visit_unit(inner, ax_expr="ax", df_expr="sub")
self.emit(f"ax.set_title(f'{facet_field}={{facet_val}}')")
self.pop_indent()
self.emit("fig.tight_layout()")
# --- data / transforms ---
def _emit_data(self, spec, df_var):
data = spec.get("data") or {}
if "values" in data:
self.emit(f"{df_var} = pd.DataFrame({json.dumps(data['values'])})")
elif "url" in data:
self.emit(f"{df_var} = pd.read_json({data['url']!r})")
elif "name" in data:
self.emit(f"# expecting a name-bound dataset {data['name']!r}")
self.emit(f"{df_var} = pd.DataFrame()")
else:
self.emit(f"{df_var} = pd.DataFrame() # spec has no data")
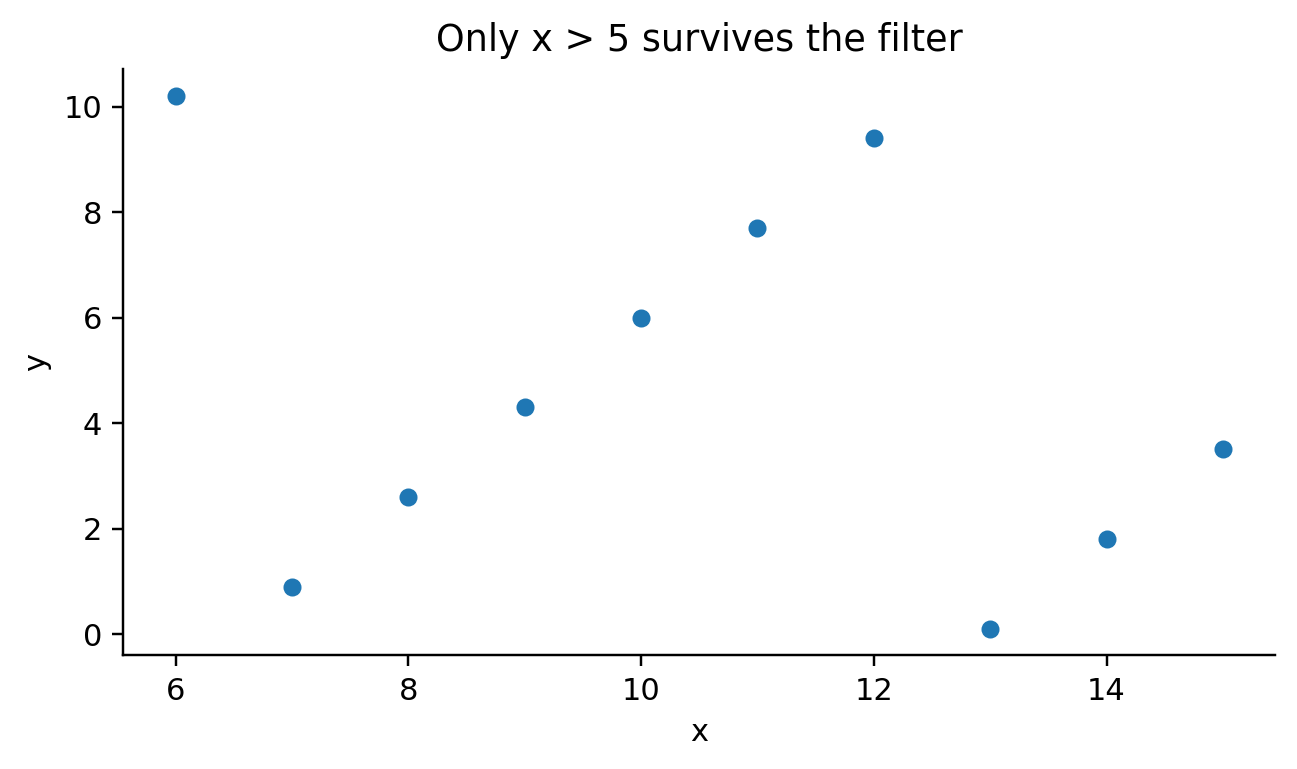
def _emit_transforms(self, spec, df_var):
for t in spec.get("transform", []):
if "filter" in t and isinstance(t["filter"], str):
py_expr = t["filter"].replace("datum.", f"{df_var}.")
self.emit(f"{df_var} = {df_var}[{py_expr}].reset_index(drop=True)")
else:
self.emit(f"# unsupported transform skipped: {t}")
# --- mark dispatch ---
def _dispatch_mark(self, spec, ax_expr, df_expr):
mark = _mark_type(spec)
method = getattr(self, f"visit_mark_{mark}", None)
if method is None:
raise NotImplementedError(f"mark {mark!r} not supported")
method(spec, ax_expr, df_expr)
# --- mark visitors ---
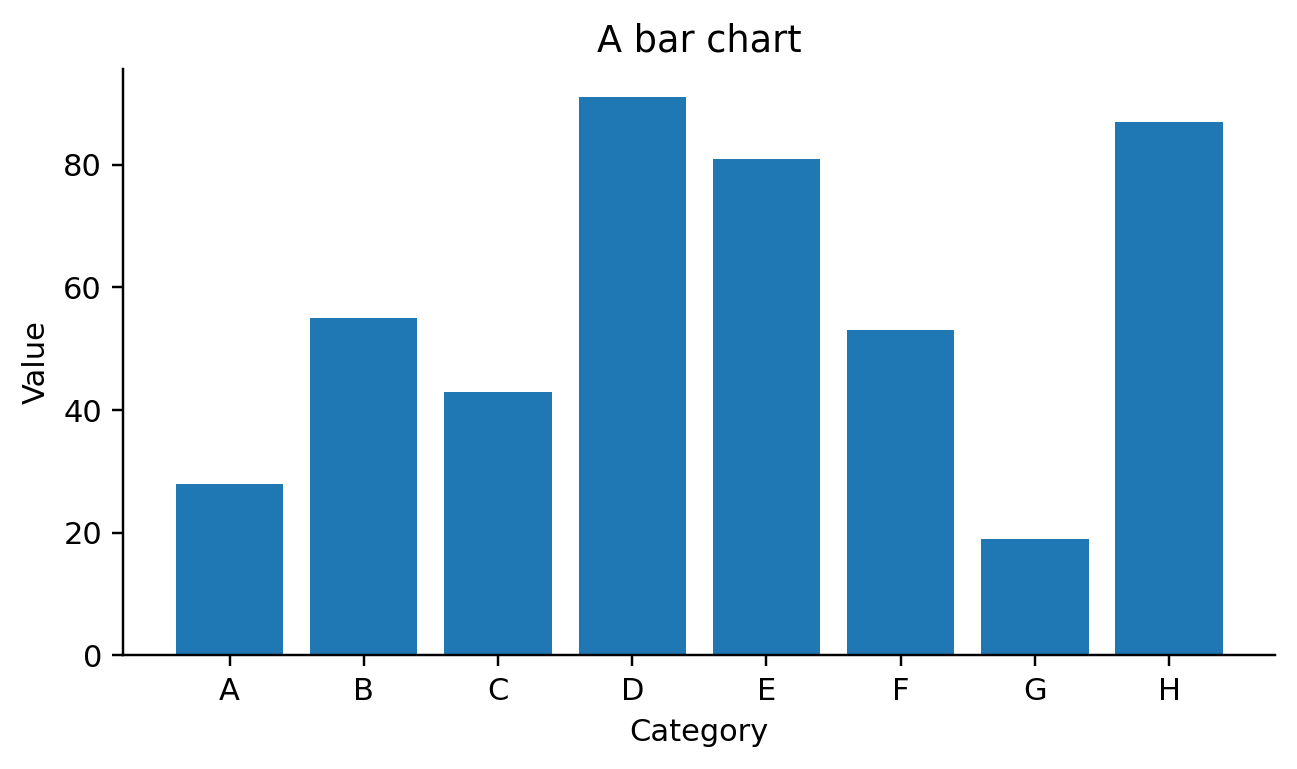
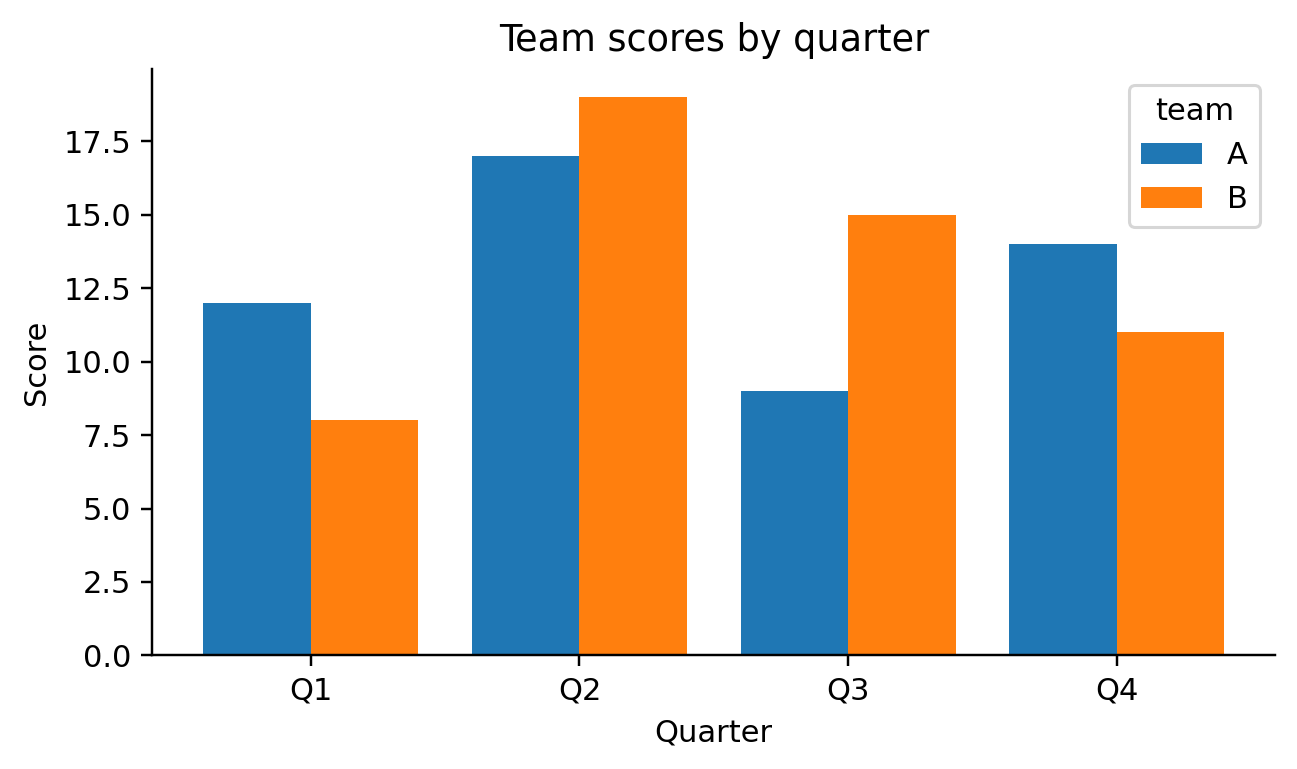
def visit_mark_bar(self, spec, ax_expr, df_expr):
enc = spec["encoding"]
x_field = enc["x"].get("field")
y_field = enc["y"].get("field")
x_agg = enc["x"].get("aggregate")
y_agg = enc["y"].get("aggregate")
color_field = enc.get("color", {}).get("field")
if y_agg in SUPPORTED_AGGREGATES and not x_agg:
self._emit_aggregated_bar(df_expr, ax_expr, x_field, y_field, y_agg,
horizontal=False, color_field=color_field)
return
if x_agg in SUPPORTED_AGGREGATES and not y_agg:
self._emit_aggregated_bar(df_expr, ax_expr, y_field, x_field, x_agg,
horizontal=True, color_field=color_field)
return
if color_field is None:
self.emit(f"{ax_expr}.bar({df_expr}[{x_field!r}], {df_expr}[{y_field!r}])")
return
self.emit(f"groups = list(dict.fromkeys({df_expr}[{color_field!r}]))")
self.emit(f"x_vals = list(dict.fromkeys({df_expr}[{x_field!r}]))")
self.emit("x_idx = np.arange(len(x_vals))")
self.emit("width = 0.8 / max(1, len(groups))")
self.emit("for i, g in enumerate(groups):")
self.push_indent()
self.emit(f"sub = {df_expr}[{df_expr}[{color_field!r}] == g]")
self.emit(f"ys = [sub[sub[{x_field!r}] == xv][{y_field!r}].sum() for xv in x_vals]")
self.emit(f"{ax_expr}.bar(x_idx + i * width - 0.4 + width / 2, ys, width, label=str(g))")
self.pop_indent()
self.emit(f"{ax_expr}.set_xticks(x_idx)")
self.emit(f"{ax_expr}.set_xticklabels(x_vals)")
self.emit(f"{ax_expr}.legend(title={color_field!r})")
def _emit_aggregated_bar(self, df_expr, ax_expr, group_field, value_field,
agg, horizontal, color_field=None):
if color_field is not None:
# group by [group, color], pivot, side-by-side bars
if agg == "count":
self.emit(
f"agg_df = ({df_expr}.groupby([{group_field!r}, {color_field!r}])"
".size().unstack(fill_value=0))"
)
else:
self.emit(
f"agg_df = ({df_expr}.groupby([{group_field!r}, {color_field!r}])"
f"[{value_field!r}].{agg}().unstack(fill_value=0))"
)
self.emit("groups = list(agg_df.columns)")
self.emit("x_vals = list(agg_df.index)")
self.emit("x_idx = np.arange(len(x_vals))")
self.emit("width = 0.8 / max(1, len(groups))")
self.emit("for i, g in enumerate(groups):")
self.push_indent()
offset = "x_idx + i * width - 0.4 + width / 2"
if horizontal:
self.emit(f"{ax_expr}.barh({offset}, agg_df[g], width, label=str(g))")
else:
self.emit(f"{ax_expr}.bar({offset}, agg_df[g], width, label=str(g))")
self.pop_indent()
if horizontal:
self.emit(f"{ax_expr}.set_yticks(x_idx)")
self.emit(f"{ax_expr}.set_yticklabels(x_vals)")
else:
self.emit(f"{ax_expr}.set_xticks(x_idx)")
self.emit(f"{ax_expr}.set_xticklabels(x_vals)")
self.emit(f"{ax_expr}.legend(title={color_field!r})")
return
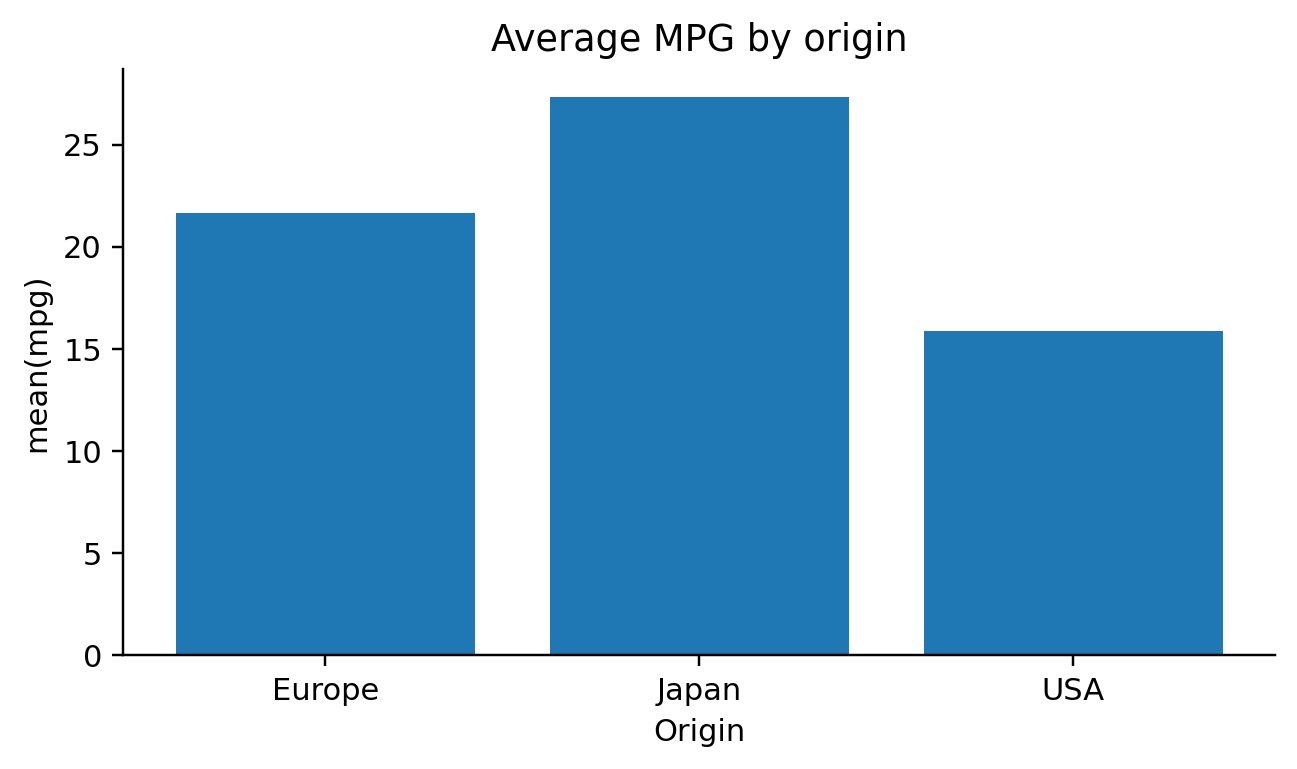
if agg == "count":
self.emit(f"agg_df = {df_expr}.groupby({group_field!r}).size().reset_index(name='value')")
value_col = "'value'"
else:
self.emit(
f"agg_df = {df_expr}.groupby({group_field!r})[{value_field!r}]"
f".{agg}().reset_index()"
)
value_col = f"{value_field!r}"
if horizontal:
self.emit(f"{ax_expr}.barh(agg_df[{group_field!r}], agg_df[{value_col}])")
else:
self.emit(f"{ax_expr}.bar(agg_df[{group_field!r}], agg_df[{value_col}])")
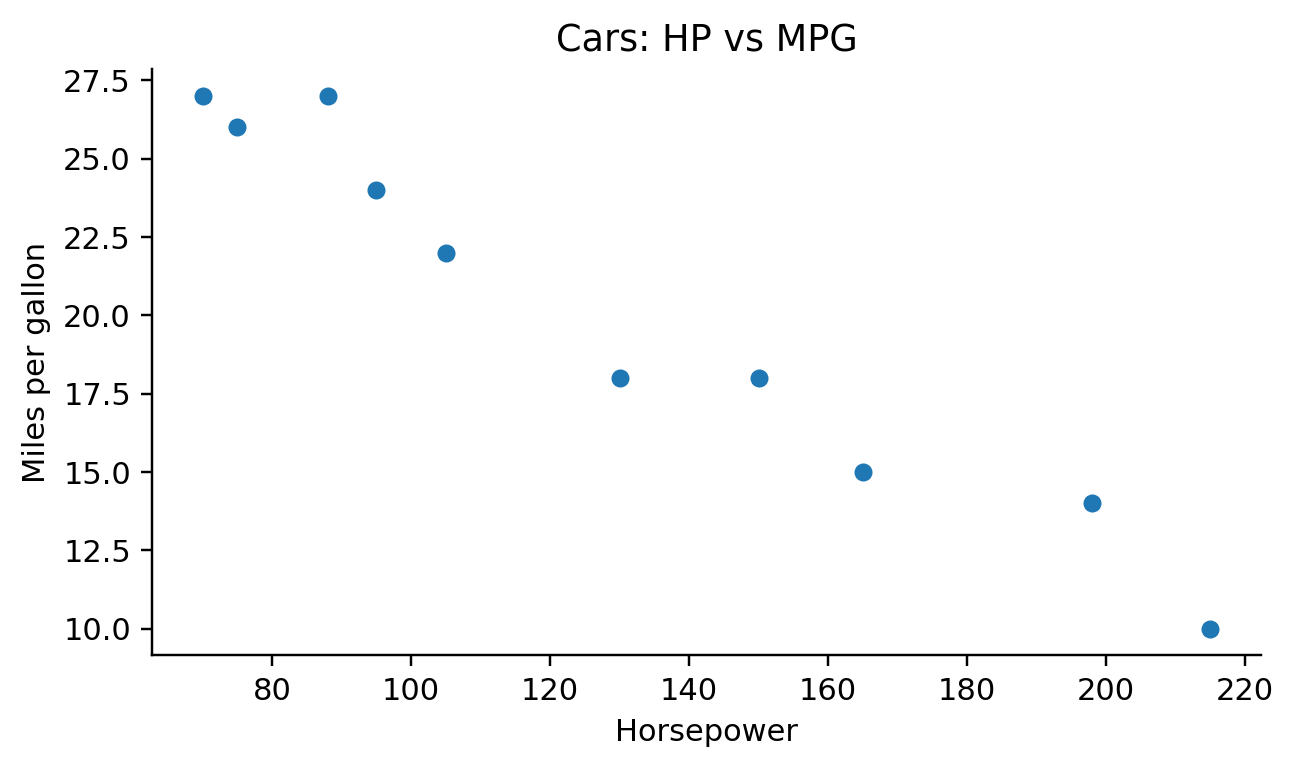
def visit_mark_point(self, spec, ax_expr, df_expr):
return self._emit_scatter(spec, ax_expr, df_expr)
def visit_mark_circle(self, spec, ax_expr, df_expr):
return self._emit_scatter(spec, ax_expr, df_expr)
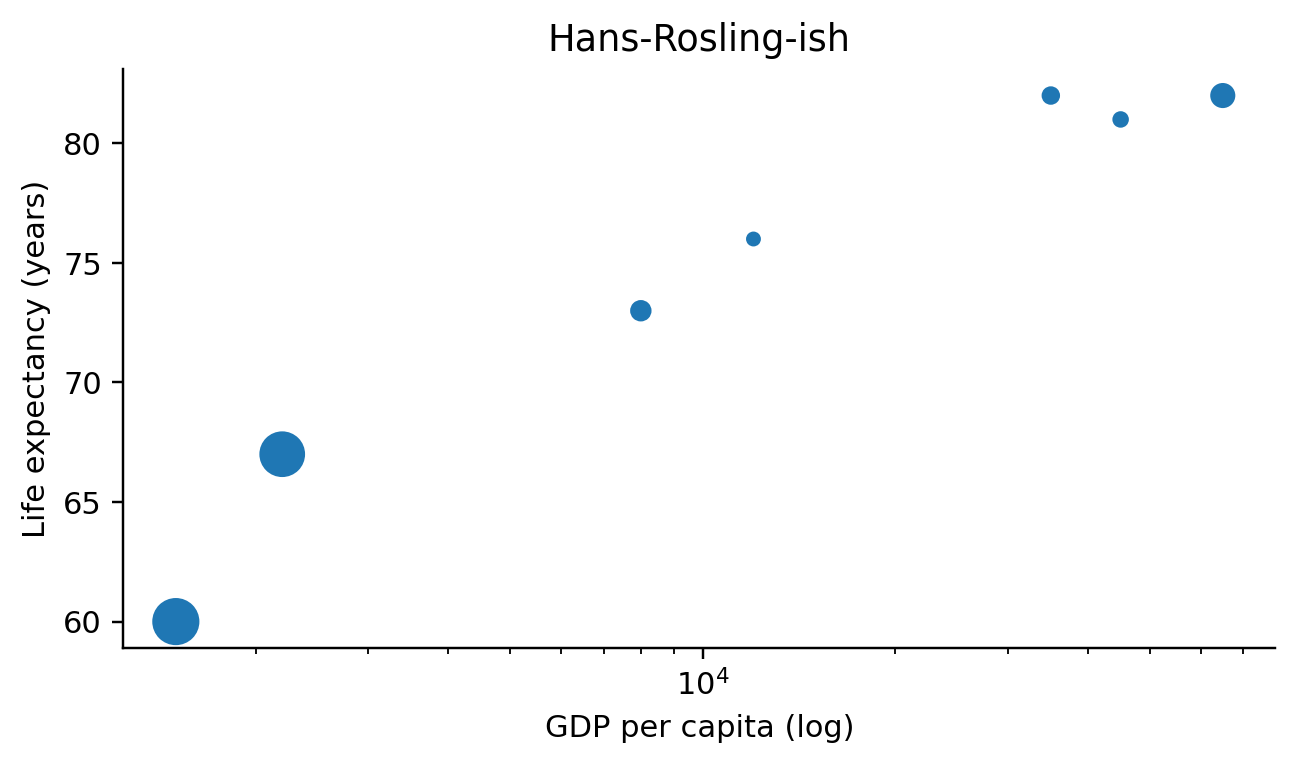
def _emit_scatter(self, spec, ax_expr, df_expr):
enc = spec.get("encoding", {})
x_field = enc["x"]["field"]
y_field = enc["y"]["field"]
color_field = enc.get("color", {}).get("field")
size_field = enc.get("size", {}).get("field")
opacity_field = enc.get("opacity", {}).get("field")
size_arg = ", s=24"
if size_field:
size_arg = (
f", s=({df_expr}[{size_field!r}] / max(1.0, {df_expr}[{size_field!r}]"
".max())) * 200 + 8"
)
alpha_arg = ""
if opacity_field:
alpha_arg = (
f", alpha=({df_expr}[{opacity_field!r}] / max(1.0, {df_expr}"
f"[{opacity_field!r}].max())).clip(0, 1)"
)
if color_field is None:
self.emit(
f"{ax_expr}.scatter({df_expr}[{x_field!r}], {df_expr}[{y_field!r}]"
f"{size_arg}{alpha_arg})"
)
return
sub_size = ", s=24"
if size_field:
sub_size = (
f", s=(sub[{size_field!r}] / max(1.0, {df_expr}[{size_field!r}]"
".max())) * 200 + 8"
)
sub_alpha = ""
if opacity_field:
sub_alpha = (
f", alpha=(sub[{opacity_field!r}] / max(1.0, {df_expr}"
f"[{opacity_field!r}].max())).clip(0, 1)"
)
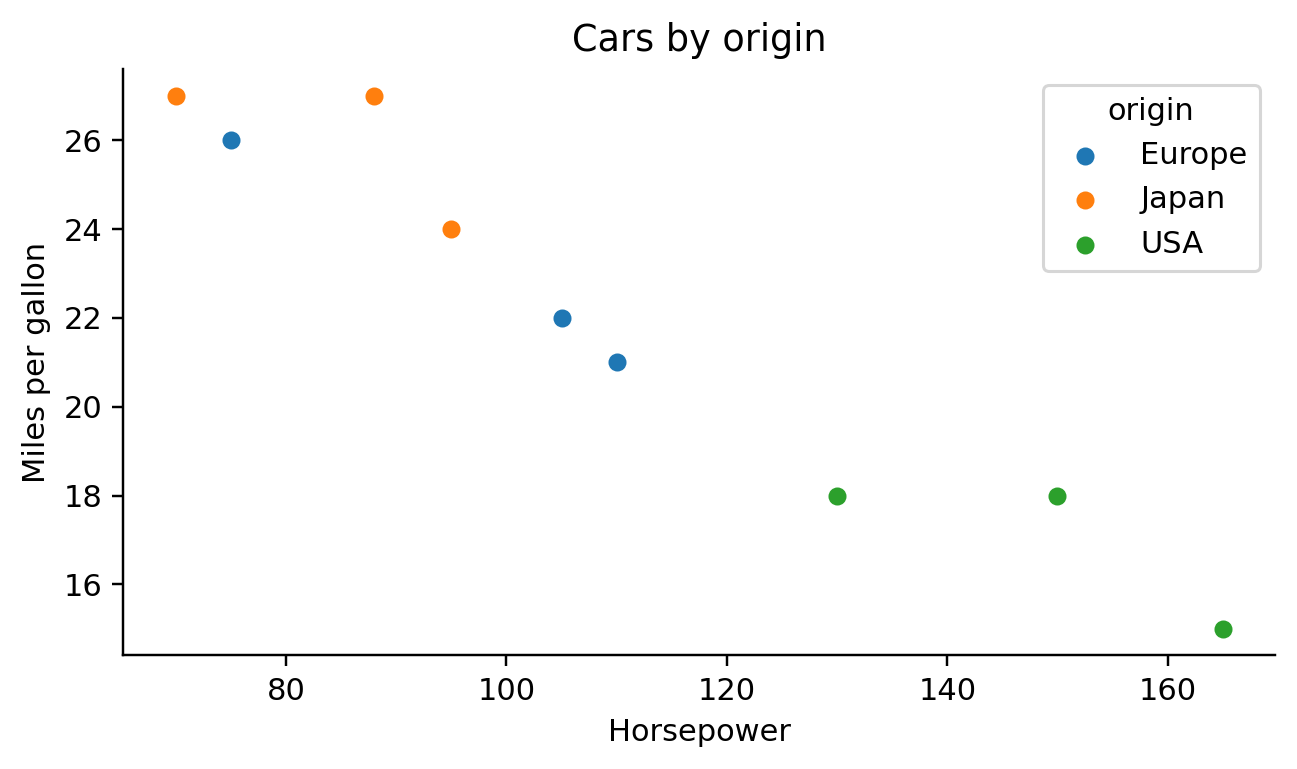
self.emit(f"for label, sub in {df_expr}.groupby({color_field!r}):")
self.push_indent()
self.emit(
f"{ax_expr}.scatter(sub[{x_field!r}], sub[{y_field!r}]"
f"{sub_size}{sub_alpha}, label=label)"
)
self.pop_indent()
self.emit(f"{ax_expr}.legend(title={color_field!r})")
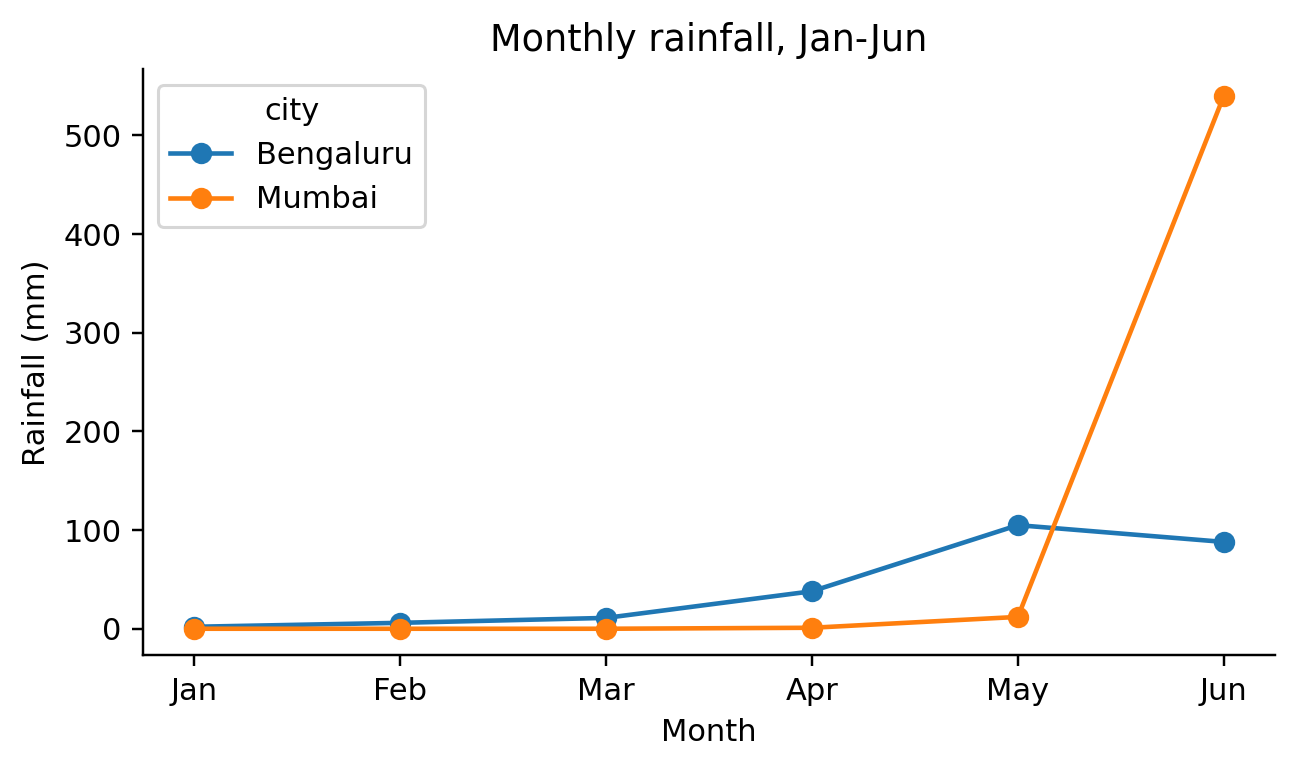
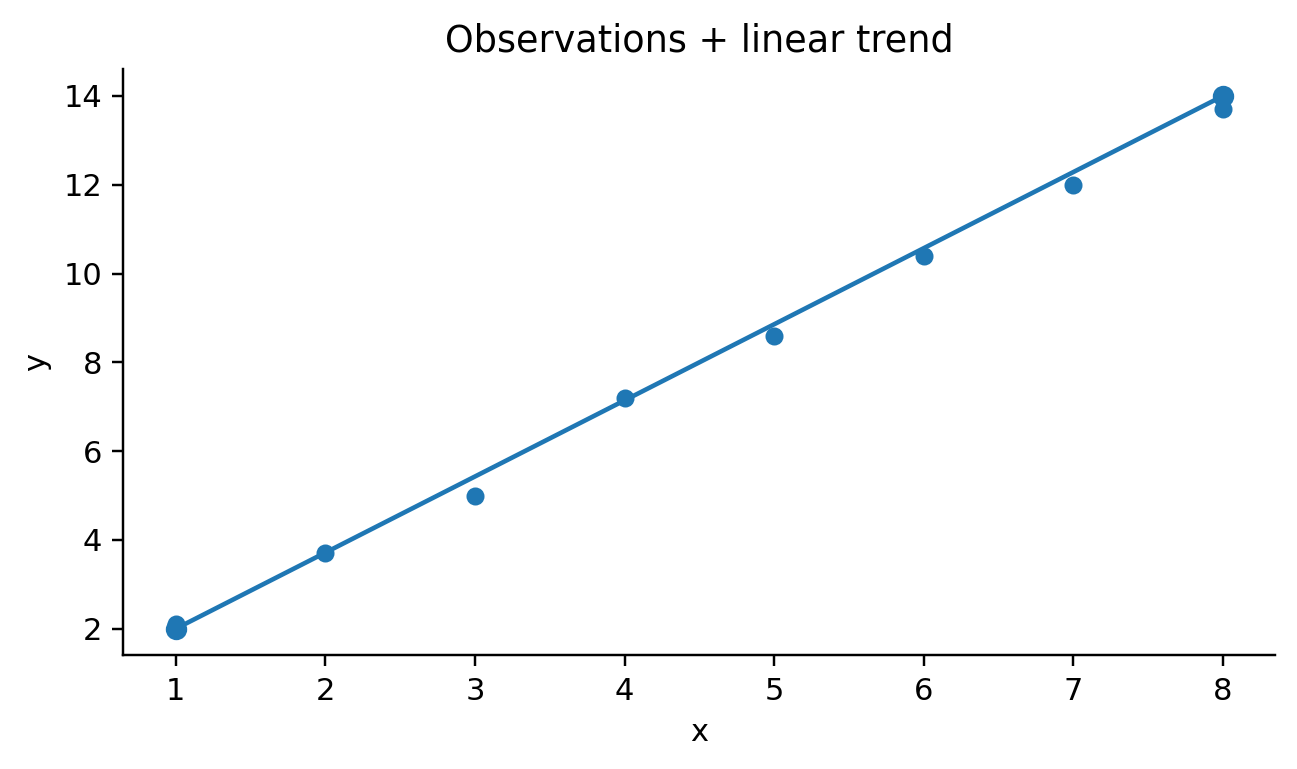
def visit_mark_line(self, spec, ax_expr, df_expr):
enc = spec.get("encoding", {})
x_field = enc["x"]["field"]
y_field = enc["y"]["field"]
color_field = enc.get("color", {}).get("field")
if color_field is None:
self.emit(
f"{ax_expr}.plot({df_expr}[{x_field!r}], {df_expr}[{y_field!r}], marker='o')"
)
return
self.emit(f"for label, sub in {df_expr}.groupby({color_field!r}):")
self.push_indent()
self.emit(
f"{ax_expr}.plot(sub[{x_field!r}], sub[{y_field!r}], marker='o', label=label)"
)
self.pop_indent()
self.emit(f"{ax_expr}.legend(title={color_field!r})")
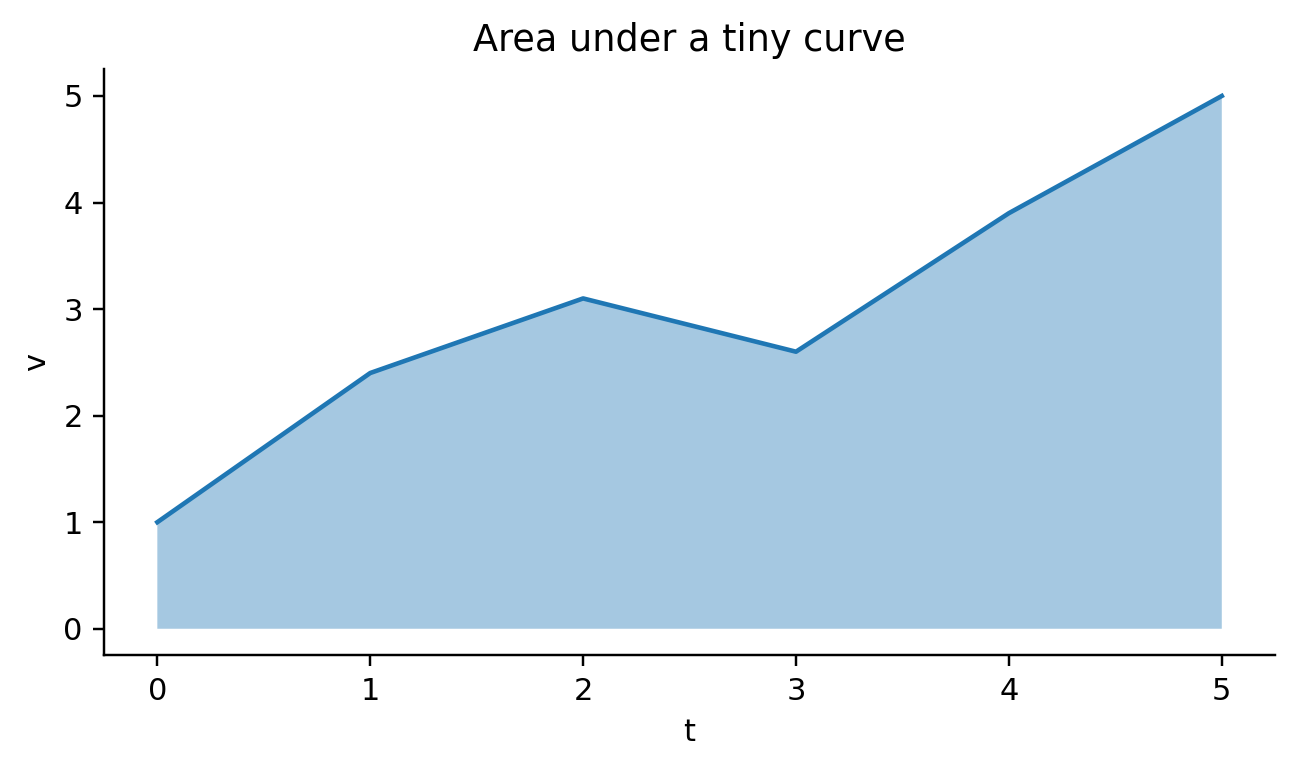
def visit_mark_area(self, spec, ax_expr, df_expr):
enc = spec.get("encoding", {})
x_field = enc["x"]["field"]
y_field = enc["y"]["field"]
color_field = enc.get("color", {}).get("field")
if color_field is None:
self.emit(
f"{ax_expr}.fill_between({df_expr}[{x_field!r}], "
f"{df_expr}[{y_field!r}], alpha=0.4)"
)
self.emit(
f"{ax_expr}.plot({df_expr}[{x_field!r}], {df_expr}[{y_field!r}])"
)
return
self.emit(f"for label, sub in {df_expr}.groupby({color_field!r}):")
self.push_indent()
self.emit(
f"{ax_expr}.fill_between(sub[{x_field!r}], sub[{y_field!r}], "
"alpha=0.35, label=label)"
)
self.pop_indent()
self.emit(f"{ax_expr}.legend(title={color_field!r})")
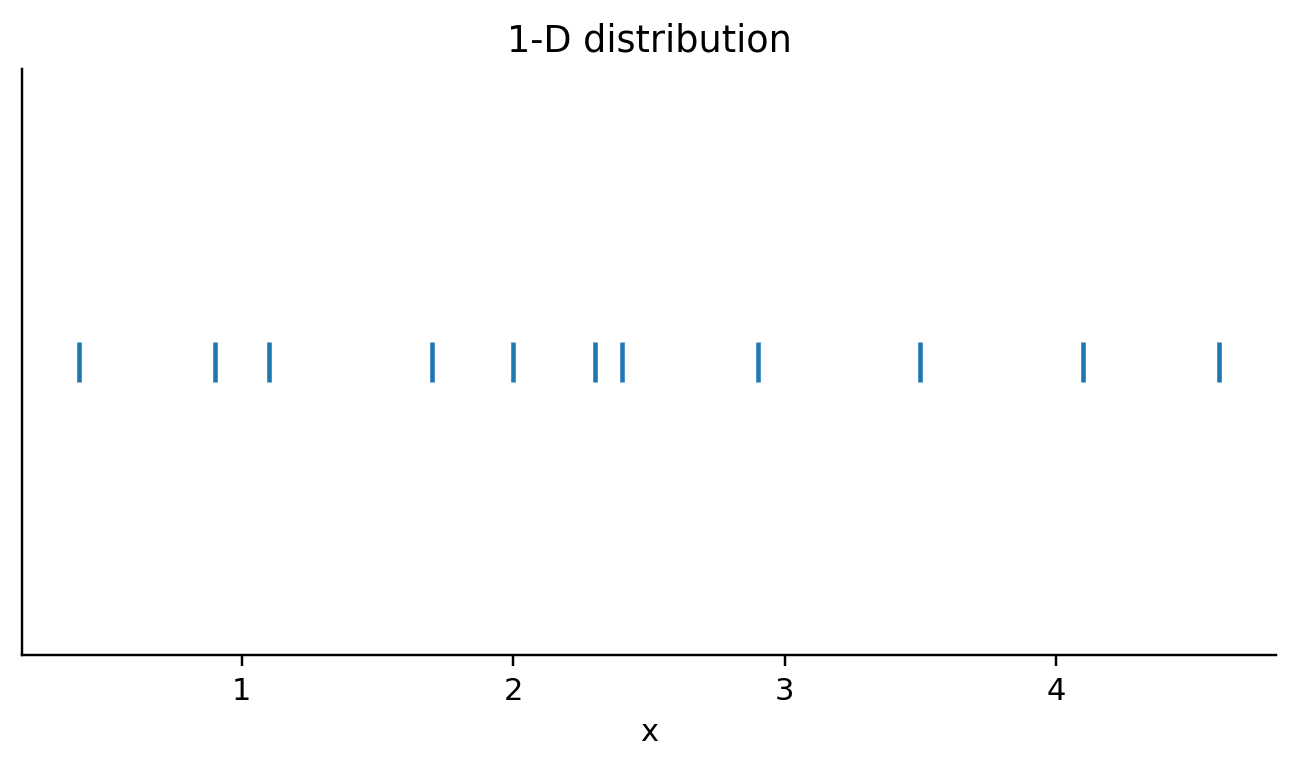
def visit_mark_tick(self, spec, ax_expr, df_expr):
enc = spec.get("encoding", {})
x_field = enc["x"]["field"]
y_field = enc.get("y", {}).get("field")
if y_field:
self.emit(
f"{ax_expr}.scatter({df_expr}[{x_field!r}], {df_expr}[{y_field!r}], "
"marker='|', s=180)"
)
else:
self.emit(
f"{ax_expr}.scatter({df_expr}[{x_field!r}], "
f"np.zeros(len({df_expr})), marker='|', s=180)"
)
self.emit(f"{ax_expr}.set_yticks([])")
# --- axis decoration ---
def _emit_axes(self, spec, ax_expr):
enc = spec.get("encoding", {})
if "x" in enc:
self.emit(f"{ax_expr}.set_xlabel({_enc_label(enc['x'])!r})")
scale = enc["x"].get("scale", {}) or {}
if scale.get("type") == "log":
self.emit(f"{ax_expr}.set_xscale('log')")
if "domain" in scale:
lo, hi = scale["domain"]
self.emit(f"{ax_expr}.set_xlim({lo}, {hi})")
if "y" in enc:
self.emit(f"{ax_expr}.set_ylabel({_enc_label(enc['y'])!r})")
scale = enc["y"].get("scale", {}) or {}
if scale.get("type") == "log":
self.emit(f"{ax_expr}.set_yscale('log')")
if "domain" in scale:
lo, hi = scale["domain"]
self.emit(f"{ax_expr}.set_ylim({lo}, {hi})")
if spec.get("title"):
self.emit(f"{ax_expr}.set_title({_unwrap_title(spec['title'])!r})")
def _unwrap_title(title):
if isinstance(title, dict):
return title.get("text", "")
return title
def vegalite_to_matplotlib(spec):
"""Backward-compatible wrapper around the visitor."""
return VegaLiteToMatplotlib(spec).code()
def render_generated(code):
exec(compile(code, "<generated-mpl>", "exec"), {})
def show_side_by_side(spec):
print("--- generated Matplotlib code ---")
code = vegalite_to_matplotlib(spec)
print(code)
print()
print("--- Altair render of the original spec ---")
display(alt.Chart.from_dict(spec).properties(width=320, height=200))
print("--- Matplotlib render of the generated code ---")
render_generated(code)